search this blog
Wednesday, October 8, 2014
Analysis of an ancient genome from Hinxton
I've just added an ancient sample from Hinxton, England, to my burgeoning ancient genomes collection. It's a pre-publication release freely available here as ERS389795. Thanks to Felix C. for breaking the news. We've both called this sample Hinxton1.
Unfortunately, its archeological context is a mystery to me, but it's possibly one of the ancient genomes mentioned in the recent Schiffels et al. ASHG abstract (see here).
In terms of genome-wide genetic structure, Hinxton1 is most similar to present-day Orcadians, Irish, western Scots, Icelanders and western Norwegians, more or less in that order. However, it's fairly distinct from the modern inhabitants of England, or at least those in my datasets, who mostly come from Kent and Cornwall.
Please note, this analysis features two different datasets: Eurogenes and Human Origins. Eurogenes, which is my own dataset, includes more populations than Human Origins, and is based on SNPs used in commercial ancestry and medical work. On the other hand, Human Origins shows a more varied sampling strategy, and is based on SNPs specifically chosen for population genetics.
Shared drift stats in the form f3(Mbuti;Hinxton1,Test) - Eurogenes dataset
Shared drift stats in the form f3(Mbuti;Hinxton1,Test) - Human Origins dataset
Eurogenes K15 4 Ancestors Oracle results
See also...
Analysis of Hinxton2 - ERS389796
Analysis of Hinxton3 - ERS389797
Analysis of Hinxton4 - ERS389798
Analysis of Hinxton5 - ERS389799
Hinxton ancient genomes roundup
Subscribe to:
Post Comments (Atom)

340 comments:
1 – 200 of 340 Newer› Newest»I was going to say Swede, then found it was in East Anglia or the Southern Danelaw. The earliest Angles had strong Swede influence, but the C-14 is a little too early.
Without knowing the context, one possibility is the man was a Roman soldier who was ethnically Angle or Swede within the Roman timeframe.
If the C-14 is correct then it could also mean that the Saxon Shore was already settled with various Scandinavians before the Roman conquests.
This may seem like a small thing to non-British people, but it could blow up in the British press.
Scandinavians were already there. He is 44% EEF, 40% WHG, 16% ANE. Very Scandinavian like. Remember the kent grave with 5 Scandinavians?
Wasn't this one supposed to be a Spanish-like Celt?
If this is indeed one of the Iron Age samples, it doesn't seem to agree with the paper's own analysis:
"We find in particular that while the Anglo-Saxon samples resemble more closely the modern British population than the earlier samples, the Iron Age samples share more low frequency variation than the later ones with present day samples from southern Europe, in particular Spain".
Interesting, assuming this guy (?) is close to the mean for his population, looks like there was still some excess Loschbour / WHG affinity in the Iron Age Celts (?) and perhaps all Northwest Europe that was reduced a little by subsequent population contacts (whether Anglo-Saxons or simple deme-to-deme flow).
When we get some more ancient samples, we can see whether Iron Age Briton has the same relative position compared to ancient Lithuanians, Spaniards, Greeks, etc. as his descendents do today - e.g. although Iron Age Briton is more Loschbour like, Iron Age Spaniards are more Oetzi like and Poles and Lithuanians are more Motala 12 like as well, etc. so all the relative position is the same. Or alternatively if some populations have shifted more than others.
You could see a population like this mixing with a Baltic population to give rise to Scandinavians, based on the PCA.
Very interesting David thanks for posting this. Both the K15 and K13 results are pretty much what I expected them to be. Would you say that the North_Sea component from the K15 calculator when seen in this old Anglo-Saxon sample represents the same type of ancestry basically when seen in modern day Europeans?
No Baltic population is needed. He's already basically a Norwegian.
He's actually most similar to the western Scots from Argyll and to some Orcadians. I haven't checked his allele sharing stats yet, but I'm betting on some solid links with Basques, considering his western position on the PCA. I'll post the full analysis later today.
Scandinavian links seem more likely. An English person, pre-saxon should logically be well more than 50% eef.
He's definitely very northern, but he's actually more Irish than Scandinavian.
I'll post some more PCA soon, based on SNPs rather than ADMIXTURE components. They show the Sottish and Irish links more clearly. I'll also compare him to the other ancient genomes.
You would think this person to be an immigrant from somewhere. His EEF is about 10% too low for modern se England. It's hard to figure another way that he is 10-20% lower than he should be.
Maybe eastern England was more WHG until the Norman invasion or even the Industrial Revolution?
Doubtful, Normans only accounted for 1% of the population when they came. A slight difference is possible, but that would be a big jump... Interesting though.
Maybe he is an elite and the locals more around 60%.
On the Norman invasion thing, the Normans were basically Scandinavians who took on French customs, culture and language. Hence the name (Northman). So would it be that Normans, (really French speaking Norwegians) were already quite high in WHG?
"You would think this person to be an immigrant from somewhere. His EEF is about 10% too low for modern se England. It's hard to figure another way that he is 10-20% lower than he should be."
If anything Germans made Britain more EEF and less WHG and ANE. This is because they had a lot of Gaulish(French-like) ancestry. Dutch and west Germans are far more French-like than Scandinavian-like.
He belonged to R1b-L51, which would be very typical of an Iron age Briton not Scandinavian.
By a quick glance this Briton seems to be an extreme-northwest European, he scores higher in northwest European centered components than Irish and Scandinavians, who score the highest today. Maybe he wasn't 100% British, Davidski will probably find out if he had Scandinavian and or Iberian ancestry.
.
R1b-L11 my bad.
We don't know what sort of impact the Normans had on English DNA, because we don't have any Norman genomes.
However, if they came from France after sitting there for a while, then I'd be surprised if they weren't part French. Also, maybe the Norman invasion facilitated migrations of common folk between England and Normandy?
But I have a feeling that what we're seeing here are genetic substructures in pre-Anglo-Saxon England. The eastern and northern part of the country was probably very Irish-like at the time, while the south and west more Cornish-like. However, after the Anglo-Saxons, Normans and the Industrial Revolution, these substructures were shifted and smoothed out, so that the Cornish are now slightly more French-like than the eastern English, who are much less Irish-like than they used to be, and more continental Germanic.
North Germany and Denmark are less EEF than England. Norman invaders were probably more French than Scandinavian, yes. Northern France and England are about identical.
Was the individual tested for anything under L11? That's what I am. I am half British, with the other half North German, Danish, Alsatian, and Jew. I am 50%EEF, 35%WHG, and 15%ANE. On the Oracle I am plotting with the SE English, at -1.0, so pretty spot on.
Oh jeez I have now realized this is not the Anglo-Saxon but the Celt. doiiii lol. His result make sense still never the less.
My mother is from southern England, here are her results for comparison:
K15
North_Sea 29.07%
Atlantic 36.46%
Baltic 7.78%
Eastern_Euro 7.86%
West_Med 11.20%
West_Asian 6.59%
East_Med -
Red_Sea 0.53%
South_Asian 0.50%
K13
North_Atlantic 51.09%
Baltic 20.33%
West_Med 15.84%
West_Asian 6.84%
East_Med 3.35%
Red_Sea 0.99%
South_Asian 1.42%
K13
North_Atlantic 49.19
Baltic 23.52
West_Med 13.43
East_Med 7.8
West_Asian 2.98
Siberian 1.44
Oceanian 0.85
Red_Sea 0.65
South_Asian 0.14
"Was the individual tested for anything under L11? That's what I am. I am half British, with the other half North German, Danish, Alsatian, and Jew. I am 50%EEF, 35%WHG, and 15%ANE. On the Oracle I am plotting with the SE English, at -1.0, so pretty spot on."
There was some arguing between members whether he has results in P312 and U106. Maybe later we'll find out what L11 clade he belonged to. BTW, he had mtDNA K1a1b1b.
He's the first ancient R1b-L11 sampled so far. It's neat that a genome of a pre-Roman west European from an ethnic group we can claim decent from who lived in a very primitive and sparse-rural world, was pagean, and had a very differnt culture compared to modern west Europeans but had the same genetic markers as us(In my case, for the most part).
You should take a Y SNP test from FTDNA, and specifically P312 and U106 and their downstream clades.
I'm l11. Negative for u106.
1/3 of the Norman Army was really Norman, 1/3 was French an Flemish and 1/3 was breton ( Brittany)
It's very interesting that an R1b is that much WHG, and not so much EEF. Either highly admixed or R1b didn't start in West Asia, but also the Steppes and is only in West Asia as Anatolian speakers, with v88 and 335 being possibly Mesolithic migrations across the Caucasus. Who the hell knows?!?
@bellbeaker
Caesar said the Belgae were German by ethnicity but Gaulish (Celtic) by culture.
http://en.wikipedia.org/wiki/Belgae
I think we're going to find those ancient writers were mostly right.
I'm still running the tests, but yeah, he's definitely a Brit, not a Scandinavian. The thing is though, his lack of Near Eastern affinity makes him look Icelandic in a couple of tests.
So if he's representative of the population that brought R1b to the British Isles, then R1b came from Hyperborea, or thereabouts.
Also, modern English are very different. But I guess that's not surprising, considering that the Romans, Anglo-Saxons, Normans, and the Industrial Revolution all came after him.
David,
That term carries lots of meanings. Are you thinking West Siberian?
No one knows where Hyperborea was exactly, kind of like the place of origin of European R1b. Hence the analogy.
Gotcha. Maybe Anthony is correct in his assessment of a steppe group splitting nw of the Black Sea, with Anatolian and the Italo-Celtic-Germanic groups splitting there. It matches with the split of R1b and R1a.
Geographically speaking. It would also explain links with the Balkans and Britains I2a split of 5-6kya.
His North Sea score is quite high. Does he fall in the so called North Sea cluster(West Norwegians/East English/Northern Dutch etc) or is he more West Scottish/Irish like?
The results so far make it look like the "Spanish-like element" of Iron Age Celts could be just minor affinity in Chromopainter.
It will be interesting to see the Anglo-Saxon sample and how the researchers arrived to their conclusion. A mixture of populations less southern than modern English can not produce the modern English, and if we assume the Norman genetic contribution was negligible many things remain a mystery.
"I'm still running the tests, but yeah, he's definitely a Brit, not a Scandinavian. The thing is though, his lack of Near Eastern affinity makes him look Icelandic in a couple of tests.
So if he's representative of the population that brought R1b to the British Isles, then R1b came from Hyperborea, or thereabouts."
Does he show genealogical connections to Irish and west Scottish, or is he just a similar mix? Do you think part of the reason he is so non-near eastern could be because Celts were not totally settled down in Britain yet, recent arrivals? Does he suggest the populations that changed west Europe after Otzi were heavy in WHG and had little ENF?
"Do you think part of the reason he is so non-near eastern could be because Celts were not totally settled down in Britain yet, recent arrivals?"
Simply from the geography it always seemed likely to me that there were two flows into Britain and Ireland: a southern one up the Atlantic coast and a northern one from Holland / Denmark with the southern one picking up more farmer dna on the way and the northern one picking up more paleo dna along the way.
I think the main thing here is evidence that some of the northern genetic flow was culturally Celtic.
So I don't think it's "Celts" so much as which Celts.
As to the reason for the change since then my personal guess would be the industrial revolution pulling people into England. For example if you look at the top 20 common surnames in England, four: Jones, Williams, Evans and Davies are either Welsh origin or disproportionately Welsh.
http://en.wikipedia.org/wiki/List_of_the_most_common_surnames_in_Europe#England
"I'm not really sure how to interpret his seemingly close relationship to western Scandinavians. This could well be a reflection of pre-Viking Norse admixture in parts of the British Isles. Alternatively, it might be due to the unusually high level of indigenous European hunter-gatherer ancestry carried by this sample, which is very obvious in almost all of the tests I was able to run."
There was significant gene flow/ contact between Britain and Scandinavia and northern Europe long before Vikings, Anglo-Saxons, et al
Even during the Bell Beaker period, which most people think of "coming from" Iberia, many of the BB artifacts found in Britain actually have closest analogies to the lower Rhine region. This has been confirmed further by suggestive isotopic analyses, eg Bronze & Iron age sites such as Cliffs End, where up to 60% of the sampled were immigrants, isotopically (!)
"Then again, perhaps it's because Icelanders and western Norwegians harbor significant Celtic ancestry?"
I understand that a significant percentage of Norwegian settlers of Iceland took with them Irish wives from their (Viking) settlements in Ireland.
@MikeThomas "Even during the Bell Beaker period, which most people think of "coming from" Iberia, many of the BB artifacts found in Britain actually have closest analogies to the lower Rhine region."
If that's the BB reflux, that population coming back out of Central Europe may have been genetically quite different from the original BB coming from Iberia into Central Europe. Even if they retained their R1b lineage, that reflux population could well have been responsible for spreading ANE across western Europe,as well as other more 'eastern' things after picking them up from Corded Ware in Central/Eastern Europe, as Davidski actually suggested a while back if I'm correct. And this guy is high in ANE for Europe.
It's tempting to link Celtic with BB,but it's also way too early for the Hallstatt/La Tene expansions. It's likely that BB was a significant migration to Britain/Ireland(isotopes), but I don't know about Hallstatt/La Tene?
It's a significant find, being only the second BC R1b after Kromsdorf, so props to Davidski and Felix C.
The EEF is quite low and the ANE quite high for any out of Iberia scenario for R1b. Iberian beaker was probably G, E, T, K, and I. R1b is quite low in NW Iberia, compared to surrounding areas.
G, E, T, and I for Iberia, sorry.
@ Shaikorth It will be interesting to see the Anglo-Saxon sample and how the researchers arrived to their conclusion. A mixture of populations less southern than modern English can not produce the modern English, and if we assume the Norman genetic contribution was negligible many things remain a mystery.
It could be that both populations (AS and Briton) were more north than the present day, and then simple increases in genetic flow just pulled everyone in Europe slightly closer together. The PCA shows that it is not by a huge degree, the Iron Age Briton is unusual in that, if we assume he is typical for his population - there are probably individuals in the Irish population who can match him, even if there's no Irish population's average that can (although maybe the Western Isles of Ireland will have a surprise).
One thing I would note though, is that, if we are likely to attribute this to large scale population movements, its not necessarily the French or the Germans here who could give downward shift. What about the Romans (albeit perhaps mostly of the Gaulish / Low Countries sort), whose demographic impact we've never been able to quantify? Like Davidski has noted, this may be before or early into the Roman occupation of Britain. The blood of the legions may help to explain matters.
Still, for levels of replacement it's a very close sample. But then so is Denmark. I wonder if this is useful in actually putting levels on how much migration would be needed, overall, from the Empire and Denmark to explain modern England.
My result K13
1 North_Sea 42.98
2 Atlantic 28.72
3 Eastern_Euro 10.07
4 Baltic 7.46
5 West_Med 6.63
6 West_Asian 1.87
7 Oceanian 1.07
Iron Age Briton.
North_Sea 43.19
Atlantic 28.88
Baltic 6.46
Eastern_Euro 11.98
West_Med 6.71
West_Asian 1.74
Red_Sea 1.01
I've favored a max 25% replacement by Saxons on several forums. There are many British named leaders of English groups, well into the Middle Ages. We know Scandinavians have been going to Britain since the early Iron Age. I don't think Romans are the only cause. This individual likely pre-dates Belgic influx from the continent as well.
It's an iron age not bronze age Briton. He's pretty recent, and was expected to be almost no different from modern Irish. So, making conclusions about the people who brought R1b-L11 to Britain based on this dude, is about as reliable as doing the same with modern Irish.
Just a little playing around here. If I use Tuscans to portray the average Roman soldier, Italian/Gaul, at 75% EEF, 13% WHG, and 12% ANE, input at 10% into the ancient sample we get 47.2% EEF, 37.3% WHG, and 15.5% ANE. Pretty close to the modern English numbers. If Belgic and Saxon input is both around 20% the numbers should remain pretty static. Continental influx during the Industrial Revolution could make up the other 3% EEF. The 3% could also be made up by Western Britains being more EEF, as well. Which is probably likely.
Davidski,
Felix translated another ancient British genome.
https://drive.google.com/folderview?id=0B7vzRsRM2aOQY1J0bDlWdVREa1E&usp=sharing&tid=0B7vzRsRM2aOQTENJUlB4OVVWeUE#list
Chad,
Did you do a Geno 2.0 test? If so you're probably Df27.
Roman-admixture in Britain, or anywhere that had strong contact with Rome is an idea people rarely consider. The U152 in Britain may not only be Celtic but also Roman. I doubt it though. You should translate using Tuscans as Roman proxies in admixture tests, I doubt English and other British will score high enough in typical west Asian components.
Barak,
I do agree with you, in a way. Looking at Tuscans, who are 13% WHG, and 12% ANE, R1b was probably as much ANE as it was WHG, if not more. Albanians are about 9%WHG and 13%ANE. It could be anything really. R1b certainly carried ANE to Western Europe.
Corded Ware is not needed to explain anything, as Northwest Europeans are only a percent or two behind the Baltic people. Corded Ware only lasted a couple hundred years before Bell Beakers took over. If it was Corded genes into Bell Beaker that spread ANE, then Western Europeans would be way lower in ANE than Eastern Europeans. That just isn't the case. It's logically impossible to come up with a scenario where the English are basically as much ANE as Poles, without a total replacement of Beaker genes with Corded genes, yet the culture and y-DNA stays the same. Bell Beaker had to be almost identical in ANE, as Corded Ware. It's a new people in Central Europe, not Iberian with a reflux. Pottery didn't change in established Beaker zones. Only weapons (daggers, armguards, arrowheads), ornamentation, and a single-grave barrow culture, in the movement West, post 2600BCE. The 7% difference between the Basque and Estonians can be explained by Mesolithic survival of ANE. I've explained that many times. What was R1b?..33/33/33, 50/25/25? It's anyone's guess. If they could pull some aDNA of the German Beakers, that would be great!
I got my y-dna from 23andme. I also ran it through Prometheus. I will do an ftdna test, at some point.
If someone has the tools to run it through a program, I can send my data to them, to check for the correct subclade.
Barak,
I doubt that I am df27. My L-11 line is Danish. Eastern Denmark and Central England are L-11 hotspots, though only 10%.
David,
Are you going to run those other samples from Felix? Thanks!
I've had a look at another of the samples, but could hardly get any SNPs to analyze.
Felix hasn't yet done any of the others. But it seems he's working on one and uploading each of the files as he goes.
This guy is from the tribe of Boudicea, right?
Dont these people belong to the type of Briton that Tacitus claimed to look "German"?
His Scandinavianess seems fitting.
BTW, I doubt that modern German genoms are a good proxy for iron age Germans. My bet is:
1. Germans recieved a massive eastward shift, that is younger than 1000 years and bases on slavic migrants from the past 1000 years.
There could also be a southward shift, because of quiet a lot French migrants (partly religious refugees)from the last 400 years.
There is also rumors that, after the 30 years war dropped the German population from 12 Million down to 6 million, Germany refilled its population by massive migration from the Balkan peninsular. Wich today I doubt, because I2a is far to low in Germany for such a scenario to be true.
Still, those stories of massive migration for compensation of population loss, dont come from nowhere.
Davidski, can you email me a zipped file of the first Iron age Britain at joesmith71440@gmail.com? Downloading it from Genetic Genealogy Tools hasn't been working.
I doubt that the person is Iceni. There is the possibility that the Iceni are actually from the Belgic migration. At least the elite. The material culture is like the other Belgic tribes of England. The location of Hinxton is actually in the region where the Catuvellani and Trinovantes territory meet. If this person is from 500BCE, they likely pre-date these tribal groups.
There's a more compact version of ERS389795 here...
https://drive.google.com/file/d/0B9o3EYTdM8lQemNTNWpGdXlRS1U/view?usp=sharing
I don't have anything to open it with. That's why I want a Zipped file. I just want to test SNPs, on online analyzers. Is the second Briton male or Female? If male could you tell what Y DNA haplogroup he has?
"I doubt that the person is Iceni. "
But why did those guys, who actually tested these corpses claim, they compared a Celt from the Iceni tribe to an Anglo Saxon? If its most likely not Icen, because its too early to be?
Felix uploaded the Y-DNA and mtDNA markers separately in much smaller files. You should be able to download those and plug them into various tools.
Or isnt this individual that Iceni with the suposed spanish (wich we dont see in these results) connections?
"Felix uploaded the Y-DNA and mtDNA markers separately in much smaller files. You should be able to download those and plug them into various tools."
The downloading isn't working but a zipped email would, and I don't know why. It's alright if you don't want to send the zipped file. Felix listed the Y and mt SNP calls, so I already got that info.
I wanted to see the Iron age Briton's pigmentation, personality-associated, and Lactose, and maybe other SNP results. I also really wanted to see personality associated SNP results of the stone age genomes.
Hinxton is a ways from Iceni territory. Maybe they aren't keen on history?
Graham Little, are you from Scotland?
Barak,
Where is Felix posting the info?
Here..
http://www.y-str.org/2014/10/hinxton-dna.html
In regards to the Spanish connection, I didn't look at the sharing of low frequency alleles between this genome and present-day Europeans. And low frequency SNP sharing isn't the same thing as overall ancestry.
But if you look at the Human Origins shared drift stats I posted above, the Basques and Pais Vasco Spaniards are actually quite high up the list.
Chad,
Hinxton is in South Cambridgeshire.
Looking at various maps, this area was either inhabited by the Iceni or the Catuvellauni, or maybe both.
I would expect anyone with a good amount of WHG and R1b to have good matches. Whether or not one comes from the other or both Spaniards and Brits received input from a similar R1b source, is the question.
Barak,
Do you have the tools to run my 23andme data for yDNA subclades?
Iceni territory ended around the Northern 1/4 of Cambridgeshire. I haven't seen a map that places the Iceni as far as Hinxton. Hinxton is on the Southern border, basically on the boundary of Catuvellauni and Trinovantes territories, according to almost all maps.
Barak,
I used the tool on his site. So far I am just L11- L151, like this individual. I am looking for anything under it now.
Barak,
Check that. Positive for L11, L151, L52, P310, P311. Negative for U106 still.
23andme I'm pretty sure only tests deep deep subclades of L11. You'll have to do another test. It's safe to assume you probably have some type of P312. I'm going to do the same, I know I have P312 and I'm pretty sure I have Df27, but I want to be 100% sure.
23andme and Geno 2.0 are like 10 years behind genetics knowledge. It really ticks me off how they keep telling people romanticized, agenda driven, simplified bull shit(Spencer Wells is a joke!). People who know what their talking about like Riech don't get their message across to the public as much, which is sad.
FTDNA is much better because it doesn't give big theories about ancient origins and has more of a variety of tests.
@Chad
"Hinxton is a ways from Iceni territory. Maybe they aren't keen on history?"
It's in what was Belgae territory regardless of which sub-tribe.
.
"This individual likely pre-dates Belgic influx from the continent as well."
The label doesn't matter to the bigger picture which is Hinxton is in what is now called East Anglia i.e. it's the exact same area that was first settled by later waves of A-S before they pushed on over the rest of the country.
So if the Angles and Belgae came via that route then an earlier group may have done so also - or just Belgae before they were called Belgae.
Caesar says the Belgae crossed the Rhine "early" to settle in the area he knew as Belgica. He didn't say when they crossed the Rhine as he didn't know - or if an earlier group crossed the Rhine and settled there before the Belgae. It's the existence of a repeated northerly route that is interesting.
I think there was a southerly "Vascones" like flow into Britain represented most nowadays by Wales and Cornwall and a more northerly Belgic / Saxon / Danish / Norman flow who were genetically similar to each other but culturally distinct due to the c. 600 year gaps between waves and they pushed the Vascones-like population back to the west - with a rebound later when the survivors immigrated back to England.
(Just from observation I think the Irish will on average turn out to have more of this Belgic-like flow and less of the Vascones-like element than the Welsh/Cornish although it will vary by region.)
#
@Fanty
"This guy is from the tribe of Boudicea, right?"
It's definitely Belgae territory.
Geneticker's admixture results for two other Iron age Britons are very southern. So, maybe the abstract wasn't talking about this one when they said Iron age Britons possibly had some ancestry from Spain.
Yes, lots of Iberian for the two females.... but not the R1b male. It's very small for him. 1% to be exact... R1b is not Iberian...
Hold on, is Genetiker claiming that the individual we believe is R1b-L11 is ~1% Gedrosia and not an R1b individual, or is he referring to a different male from the same site?
Gedrosia must simply be a mix of Near Eastern and ANE, that is made by mixing of Near Eastern like Iberians or Europeans and ANE heavy R1b
Kind of what I was saying. Iberian Beaker is female movement.. not males.
Geneticker is very stubborn and sticks to theories he creates by looking at data for a few days(or hours). He stubbornly believes no one can belong to R1b and not score in Gedorisa. The first Iron age Briton belonged to R1b-L11, end of story.
"Kind of what I was saying. Iberian Beaker is female movement.. not males. "
This is all Iron age. Them being female is probably random. It's interesting that the girls were probably immigrants or daughters of immigrants(they weren't sisters, they had different mtDNA) from Spain or southwest France(They may have spoke ancestral form of Basque).
Britain's isolation in ancient times is maybe sometimes exaggerated. The Romans made a big deal about the isolation of Britain, and Josephus(famous ancient Jew) said they were unknown by the world till Rome discovered them.
If people all the way from Spain(maybe the girl's grandparents came from France) sailed there that's pretty amazing, it means there was direct contact or close to direct between Spain and Britain. Iberian, Roman, Gaulish, and pre-Viking Scandinavian ancestry has to be seriously considered for modern British.
These Iron age samples may be from Roman times or at least when Iberia was Roman territory, so maybe Roman-Spaniards immigrated to Roman-Britain.
I'm referring to the fact that some try to put R1b in Iberia in 3000BCE and migrating back to Central Europe with Bell Beaker. This obviously did not happen if the R1b male is only 1% Iberian. Well under modern Brits.
There were 2 Iberians in that Kent, Iron Age grave which also had 5 Scandinavians. They probably are recent with those scores.. I'm interested in the R1b male with next to no Iberian ancestry.
Genetiker's analyses are really basic and just awful.
There's something wrong with his thought process too.
Are you saying you have info from the site? The two girls don't look 100% Iberian, but 50% British and 50% Iberian. If the R1b-L11 dude is listed as one of the Scandinavian ones, he should also be 50% British. Why the hell would the authors pick these samples if they were immigrants, not representative of the ancient Britons?
Can you confirm the Iberianism of the two other Britons?
I'll get those files and run them this weekend. Till then I wouldn't get too excited about what Genetiker says about them.
R1b-L11 male
MDLP K=12
•26.46% Celto-Germanic
•22.97% East-European
•20.22% Balto-Finnic
•17.82% Paleo-Mediterranean
•3.76% Alatic-Turkic
•2.77% Uralic-Permic
•2.73% South-Central Asian
•1.73% Paleo-Balkanic
•0.85% Iberian
•0.68% Paleo-North-European
•0.01% Caucasian
•0.00% Volga-Uralic
1st Female
MDLP K=12
•40.53% Celto-Germanic
•15.52% Iberian
•15.21% Paleo-Mediterranean
•11.27% East-European
•6.69% Volga-Uralic
•6.21% Caucasian
•2.93% South-Central Asian
•1.63% Paleo-North-European
•0.01% Balto-Finnic
•0.00% Alatic-Turkic
•0.00% Paleo-Balkanic
•0.00% Uralic-Permic
2nd Female
MDLP K=12
•38.93% Celto-Germanic
•30.87% Iberian
•8.53% Caucasian
•6.04% Volga-Uralic
•5.59% South-Central Asian
•3.52% Paleo-Balkanic
•3.37% Alatic-Turkic
•2.00% East-European
•1.15% Paleo-North-European
•0.00% Balto-Finnic
•0.00% Paleo-Mediterranean
•0.00% Uralic-Permic
R1b male
MDLP K=10
•35.25% East-European
•28.32% British
•19.17% Paleo-Mediterranean
•5.73% Altaic-Turkic
•4.50% Paleo-North-European
•3.31% South-Central-Asian
•1.97% Paleo-Balkanic
•1.71% Iberian
•0.02% Caucasian
•0.01% Volga-Finnic
1st Female
MDLP K=10
•41.00% British
•17.14% Iberian
•15.50% Paleo-Mediterranean
•12.61% East-European
•7.49% Caucasian
•4.20% South-Central-Asian
•1.45% Volga-Finnic
•0.61% Paleo-North-European
•0.00% Altaic-Turkic
•0.00% Paleo-Balkanic
2nd Female
MDLP K=10
•38.80% British
•32.24% Iberian
•9.92% Caucasian
•5.61% South-Central-Asian
•4.44% Volga-Finnic
•4.11% Altaic-Turkic
•3.54% Paleo-Balkanic
•1.02% East-European
•0.30% Paleo-North-European
•0.00% Paleo-Mediterranean
Davidski,
Would you say Germans are a mix of French, Scandinavian, and Polish-like ancestors? Do Germans and especially Austrians have any Italian and or Balkan-like admixture? Do Swiss have any Italian-like admixture? Would you say Polish+Balkan-like ancestry in Austrians and east Germans is higher than Scandnavian+French-like ancestry?
I'm just wondering about my own ancestry. Some family members 23andme results are coming in.
Where's a link to that test where you take people's raw data and break down their European ancestry?
Well, you should be able to work out all of that with the Eurogenes tools and blog articles.
I'm not analyzing data at the moment, but anyway, I've just updated the ancient genomes spreadsheet with the results of the Iron Age Briton.
http://bga101.blogspot.com.au/2014/07/model-yourself-as-mixture-of-ancient.html
Awesome. Thanks, David. I'm looking rather Iron Age British.
Using 1 population approximation:
1 Iron_Age_Briton @ 8.672021
2 Ajvide70 @ 24.346138
3 Ajvide58 @ 25.372384
4 La_Brana-1 @ 28.153936
5 Loschbour @ 29.307089
6 Motala12 @ 31.723903
7 StoraFörvar11 @ 37.826317
8 Gokhem7 @ 39.181306
9 Gokhem2 @ 43.651765
10 AG-2 @ 50.628251
11 Gokhem4 @ 55.785831
12 Sardinian @ 56.752717
13 Oetzi @ 57.22691
14 Lezgin @ 57.690571
15 Stuttgart @ 63.304915
16 MA-1 @ 63.609094
17 Yemenite_Jewish @ 78.545077
18 Australian_Aboriginal @ 87.385829
19 Ethiopian_Ari_cultivator @ 93.659656
20 Sakilli @ 93.700963
25 iterations.
ANE K7 results for the Iron Age Briton
ANE 16.04
ASE 0.02
WHG-UHG 66.95
East_Eurasian 0
West_African 0.23
East_African 0
ENF 16.76
The lack of noise is great to see. The SNPs used must be those with among the highest coverage.
Nice. Cheers. :)
Using 1 population approximation:
1 Iron_Age_Briton @ 2.941211
2 Ajvide70 @ 27.453387
3 Ajvide58 @ 27.748502
4 La_Brana-1 @ 27.804632
5 Loschbour @ 28.429751
Using 4 populations approximation:
1 Iron_Age_Briton+Iron_Age_Briton+Iron_Age_Briton+Iron_Age_Briton @ 2.941211
2 Iron_Age_Briton+Iron_Age_Briton+Iron_Age_Briton+Loschbour @ 6.758911
3 Ajvide70+Iron_Age_Briton+Iron_Age_Briton+Iron_Age_Briton @ 6.955996
@David:
What was that about calibrating?
How? Or doesnt it work as you thought it would and thats why you deleted the comment?
I didn't delete the comment. It's in the other thread.
http://bga101.blogspot.com.au/2014/07/model-yourself-as-mixture-of-ancient.html
Here's my results for comparison. I am wondering if almost all Iberian ancestry is from the Iron Age. If it was there before that, you would expect the R1b male to have some. Female exogamy during metal trade???
MDLP K10
# Population Percent
1 British 38.17
2 East_European 20.93
3 Iberian 15.02
4 Paleo_Mediterranean 14.81
5 Caucasian 3.74
6 Paleo_North_European 3.46
7 South_Central_Asian 1.91
8 Paleo_Balkanic 1.28
MDLP K12
1 Celto_Germanic 36.53
2 East_European 17.72
3 Paleo_Mediterranean 14.12
4 Iberian 13.95
5 Balto_Finnic 8.46
6 Caucasian 3.12
7 Paleo_North_European 1.95
8 South_Central_Asian 1.77
9 Paleo_Balkanic 1.21
One of the Iron Age women from Hinxton was wearing a Roman ring.
Maybe the guy was a Belgae Celt and the two women were of Roman origin?
http://journals.cambridge.org/action/displayAbstract?fromPage=online&aid=8892531&fileId=S0079497X00002012
I am wondering if these women are descended from Gallic tribes that recently arrived, and the male a Belgae....The first woman is essentially the same as me and basically modern SE English... very interesting...Their Med isn't too high and the Celto-Germanic is still up there.. so more North than Italy..
The second one could have a Roman father of Iberian origin, or something to that effect...
It's difficult to say much based on those results because of the calculator effect.
Genetiker has given results for all samples now and claims ERS389795, 389796, and 389799 are the Anglo-Saxons and that 389797 and 389798 are the Celts.
He cites the former's Baltic affinity and the latter's Iberian affinity as evidence.
If this is true, it is strange to see an Anglo-Saxon so close to the Scots, Irish and Orcadians...
Shit, these are hard to decifer.. the male looks Eastern European mixed with North German, then next more NW and further into Eastern Europe. The numbers are kind of all over, looking almost like a pan Northern Europe mutt...
'One of the Iron Age women from Hinxton was wearing a Roman ring. Maybe the guy was a Belgae Celt and the two women were of Roman origin?'
The ring was probably a status symbol rather than a signifier that she's Roman.
I think it's a little more complicated than Genetiker is claiming. Southern Britain and Northern Britain are a good ways apart in EEF. Also, if this Iberian was pan-Celtic, it should be in the North too. Or we have a complete population replacement in the south with Gallic and Belgic tribes...
The Saxon thing, is possible though.. L11 is more about the Danish and Mercia, which has a lot of Anglo-Saxon ancestry. I would say that they are probably Angle, given the location and L-11.
ERS389795 is a large file with a lot of high quality markers. I'd be surprised if this wasn't the 12x read depth sequence from the Iron Age.
Also, the Hinxton Iron Age burial site had three people, not two. So there can only be two Anglo-Saxon genomes in this collection, not three.
From the abstract:
"two of which are dated to around 2,000 years before present (Iron Age), and three to around 1,300 years before present (Anglo-Saxon period)"
Genetiker's calculator results look like shit, and they can't be compared to the population averages because of the calculator effect.
I'll have to get all of those files and then see how they come out using PCA and the K13 and K15.
Hmm.. This would also mean a big population turnover in Denmark and Northern Germany. Danes must've been quite a bit more Baltic than the Angles, Saxons, and Jutes.
This is indeed possible, because Danes came from Skane.
Could that make one woman a Celt and the other Half Anglo-Saxon? She looks like she could be with the Eastern markers and the Baltic. Only half the Iberian too. Basically a modern SE English, who would basically be 50% Anglo-Saxon, if it all translates that way. A rather simplistic view, but it adds up.
Well, this is from Wikipedia...
"Historians believe that before the arrival of the precursors to the Danes, who came from the east Danish islands (Zealand) and Skåne and spoke an early form of North Germanic, most of Jutland and the nearest islands were settled by Jutes. They were later invited to Great Britain as mercenaries by Brythonic King Vortigern and were granted the south-eastern territories of Kent, the Isle of Wight among other areas, where they settled. They were later absorbed or ethnically cleansed by the invading Angles and Saxons, who formed the Anglo-Saxons. The remaining population in Jutland assimilated in with the Danes."
http://en.wikipedia.org/wiki/Denmark
Probably still back to the Iberian being reserved for Southern/Gallic/Belgic Celts of Britain, since modern Scots are the same as this Anglo-Saxon. Now it makes sense why the old writers say they looked like Germans in the North, because they were essentially the same. I suppose that also the result of Denmark and Britain receiving genes from the same German Beakers too. Corded was the type of both.
Basically there is going to be no way to tell how much genetic influence the invaders had on Northern Britain without checking the sharing with modern populations, correct?
"Genetiker has given results for all samples now and claims ERS389795, 389796, and 389799 are the Anglo-Saxons and that 389797 and 389798 are the Celts."
Personally I think the Celt vs German thing may be looking at it the wrong way.
I think the genetic groups are likely to turn out to be the elements like "Baltic", "Atlantic", "North Sea" etc and although the cultural layers will often correlate they won't always due to the way (judging by later histories) individual tribes combined into federations so an ethnically "Germanic" tribe that joined a "Celtic" federation e.g. by being invited over as mercenaries in a civil war, could over time adopt the Celtic cultural elements.
So if those markers are correct then I think they'll be "Baltic" or "Atlantic" type markers (or whatever).
Genetiker's analyses seem wrong - when you talk about the calculator effect, I guess this means that you've run the Hinxton sample within your normal ADMIXTURE run, rather than just a calculator?
Also just checking, but this sample (ERS389795) looks pretty normally European in a world context right - like world PCA? The Bronze Age Dane sample, M4, overlapped with Finns in a world context (presumably due to higher Mesolithic ancestry, rather than East Asian admixture in his case).
Genetiker's results suffer from the calculator effect because the population averages for the calculators he's using come from samples that were used in the original ADMIXTURE run. What this means is that the population averages are useless for samples tested with the calculators, unless maybe one can employ some sort of correction for the bias, which is actually a very similar problem to the PCA projection bias I blogged about recently.
The Eurogenes K13 and K15 population averages are based on samples not included in the original ADMIXTURE runs. What this means is that both the reference samples and test samples are tested under the same conditions, and so their results can be directly compared to each other.
I haven't run a global PCA with ERS389795 yet. I'll do that now.
Please note, I renamed the sample to Hinxton1 and edited the blog entry, because I have no idea which period it's from.
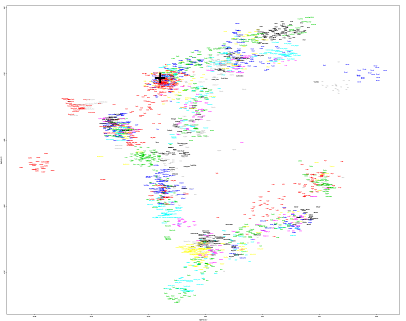
Here's that global PCA...
https://drive.google.com/file/d/0B9o3EYTdM8lQb2ljNE1IRllNaVU/view?usp=sharing
He's not easy to spot, but he's sitting with the Norwegians and Orcadians, just below the last Scot. So he's well within the range of modern Northwest Europeans, unlike the Bronze Age Dane, although more easterly than most British samples, except some Orcadians.
Felix just uploaded ERS389796, aka Hinxton2, so I grabbed it. It's very similar to Hinxton1, but with clearly higher ANE.
K13
North_Atlantic 59.1
Baltic 23.18
West_Med 8.63
West_Asian 5.87
East_Med 0.37
Red_Sea 0
South_Asian 0.93
East_Asian 0
Siberian 0
Amerindian 0.49
Oceanian 0.68
Northeast_African 0
Sub-Saharan 0.75
K15
North_Sea 42.46
Atlantic 28.9
Baltic 6.82
Eastern_Euro 13.04
West_Med 5.35
West_Asian 2.27
East_Med 0.01
Red_Sea 0
South_Asian 0.78
Southeast_Asian 0
Siberian 0
Amerindian 0.04
Oceanian 0.31
Northeast_African 0
Sub-Saharan 0.02
K7
ANE 17.52
ASE 1.06
WHG-UHG 67.2
East_Eurasian 0.33
West_African 0.55
East_African 1.28
ENF 12.07
Besides scoring higher in east Euro than northwest Europeans(but around as much as Scandinavians), these 2 Iron age Brits seem to be northwest European on steroids. It's like if French were the most southwest-type Europeans known then we got an ancient genome of a Spaniard.
How do you explain this?
I don't think they're Iron Age Brits. They're probably 2 of the 3 early Anglo-Saxons.
It seems like there might have been a population turnover on the Jutland peninsula after the Anglo-Saxon migrations to England, with the proto-Danes from the east bringing more Baltic influence.
And there was also probably a major homogenization since the medieval period across all of Northwest Europe, maybe as a result of the Great Northern wars, Industrial Revolution, and stuff like that.
Would they be good proxies for proto-Germans then? Or just the German ancestors of west and east German speakers? Do you think the northwest-European trend mostly comes from Indo Europeans, natives, or a mix of the two?
Would you explain their Scottish-Irish affinities by Scottish-Irish being at the tip of the northwest European trend and these samples having some Britonnic ancestry?
Why do you think Swedish and Norwegian are more eastern? Is it just because they turned out differently than Jutes during the Iron age, or did they used to be Jute-like and admixed with another people?
Davidski,
It makes more sense to me that they were Anglo-Saxons but it's not a huge suprise if they were Iron Age Brits either.
If these samples belonged to the Belgic tribe than it could mean that some Belgic tribes were rather Celtized and a distinct people and part of the North Sea cluster.
So i guess that modern Danes are mixture of Proto Danes originally from Scania and Jutes?
I suspect that these are good examples of early West Germanics, who were more similar to northern Celts than the more easterly North Germanics.
They basically cluster in the middle of the North Sea, if we ignore their high Icelandic affinity. Modern Danes don't cluster there because they're a mixture of North and West Germanics, and probably other people too, like Obodrite Slavs.
Of course, we need some ancient genomes of Jutes, Angles and Saxons from continental Europe to be sure of anything.
Celts and Germans spring from the same ie group, so no surprises. The nw Europe cluster is huge. I'll now bet the Bronze Age Dane is 15% EEF, 55% WHG, and 30% ANE.
Probably L11 also.
For fun I played around with Polish, Danish-Norwegian-Swedish, and French(northeastern?). It takes like 10 minutes to see you can get very similar scores to modern Germans if you mix those populations up.
I know Polish have some western ancestry, Norse have some eastern ancestry, and French may have some German ancestry and might not be good proxies of Gauls that lived in Germany, but overall these populations seem to be decent proxies for Germans ancient ancestors. West Germans don't seem to have much Polish-like ancestry at all which makes sense, but are French+Scandnavian. Scandnavian seems to be the biggest contributor for all Germans, but never over 60%.
I replaced the Norse with Hex 1 and 2, and it didn't work out at all. Hex 1 and 2 score way to low in Baltic, west Asian, and east med. And they score way to high in North Sea and Atlantic. Plus they don't score anything in most non-west Eurasian components, unlike populations on the K13 and K15 spreadsheets. That may make their North Sea and Atlantic scores look even more extreme than they are in reality compared to modern populations. It would be smart to replace modern pops non-west Eurasian scores with Hex's, for a better comparison, but that also creates problems.
"Celts and Germans spring from the same ie group, so no surprises. The nw Europe cluster is huge. I'll now bet the Bronze Age Dane is 15% EEF, 55% WHG, and 30% ANE."
I'm sure it's more complicated than that. Even in Roman times and the middle ages with hardly any writings(from mostly foreign sources), we learn that there were tons of different tribes running all over the place and mixing with each other. We don't have writings from the copper-Iron ages, so it's even harder to determine who's ancestors were who during that time, and what migrations-admixture events occurred.
Because of this it's very hard to determine exactly what any pre-modern population was like, or the ancient common origin of Iron age ethnic groups. Hex 1 and 2 kind of prove that, because they're northwest European like, but can't fit perfectly with any modern ones, and they're as recent as the Iron age or early middle ages.
Based on admixture we could also say Finno-Urgics and Balts spring from the same ethnic source. In my opinion all we can say is there's north and south trend in west Europe.
Celts in Britain and Ireland and Germans from Scandinavia were at the northern tip, Celts in France and central Europe were in the middle, and Celts-Iberians in Iberia were at the southern tip.
Celts seem to have been very diverse genetically like the Slavs, and could have spread as recently as the Iron age. How do we know if the southwest-like ancestry in Germans is only from the Gauls and not an unknown non-Celtic linguistic group?
The ASHG conference will probably reveal a lot about the Neolithic-bronze age ancestors of east Europeans, and maybe a Bell Beaker or Unetice(or whatever) genome will tell what changed west Europe after the Neolithic.
Felix published his GEDmatch analysis of Hinxton-2 today:
http://www.fc.id.au/2014/10/hinxton-2-analysis.html
So you guys can re-argue everything on the mtDNA side. :)
"I don't think they're Iron Age Brits. They're probably 2 of the 3 early Anglo-Saxons."
Yes, that makes a lot more sense than the initial speculation of being Iron Age samples. It's hard to find any Iberian traces in these as the abstract stated (and they do represent modern day Brits quite closely, if just slightly more Nordic).
We'll see if the other ones confirm or deny this.
I filtered out the duplicate SNPs from Hinxton2's file which were causing a bit of noise. Here are the new results.
K13
North_Atlantic 62.5
Baltic 25.06
West_Med 7.47
West_Asian 4.66
East_Med 0.02
Red_Sea 0
South_Asian 0
East_Asian 0
Siberian 0
Amerindian 0
Oceanian 0.29
Northeast_African 0
Sub-Saharan 0
K15
North_Sea 45.05
Atlantic 31.08
Baltic 7.2
Eastern_Euro 12.1
West_Med 4.19
West_Asian 0.4
East_Med 0
Red_Sea 0
South_Asian 0
Southeast_Asian 0
Siberian 0
Amerindian 0
Oceanian 0
Northeast_African 0
Sub-Saharan 0
K7
ANE 17.42
ASE 0.62
WHG-UHG 70.72
East_Eurasian 0
West_African 0.27
East_African 0.19
ENF 10.78
Holy shit, that's even more extreme than before! People like her were might have been the ones who changed west Europe after the Neolithic. It's an interesting idea.
Here's her Dodecade K12b score...
Gedrosia 10.69%
Siberian 0.76%
Northwest_African 0
Southeast_Asian 0
Atlantic_Med 35.86%
North_European 50.25%
South_Asian 0.11%
East_African 0.62%
Southwest_Asian 0
East_Asian 0
Caucasus 0.99%
Sub_Saharan 0.72%
Her Gedorsia is higher than any modern pops, by about 1%. I've heard Gedorsia is associated with ANE and R1b in west Europe. This could be more evidence people like her(or even more extreme) are the ones who changed west Europe after the Neolithic. If, so where did all the WHG come from?
Her Gedrosia isn't really that high. It's the calculator effect.
Hinxton2's just outside the range of modern NW Euro variation.
https://drive.google.com/file/d/0B9o3EYTdM8lQelRzYlNtR2dDbk0/view?usp=sharing
"Her Gedrosia isn't really that high. It's the calculator effect."
She scores the same in ANE K7, K13, and K15 on GEDmatch as what you got for her.
Hmmm... So these two samples (so far) might be Anglo Saxon? Seems like a strange world where early Anglo Saxons (perhaps before mixing with the people living in Britain?) might, autosomally, have been slightly exaggerated Irish / Scottish (shared drift, position on the PCA) compared to modern English, but there it is if so.
The abstract on rare allele sharing singles out the 1000GP Spain as above the 1000GP Toscani in sharing with the Iron Age samples (in particular Spain), although it seems to imply there is greater sharing for both relative to how much they share with Anglo Saxon. It'll be interesting to see if this is mostly because the Spain samples aren't as far south (genetically) as the Toscani, or if there is a much larger sharing gap than this would imply and greater sharing with Spain is more due to Spain's further west position.
Unlike the Dodecad pop averages, the K13 and K15 pop averages aren't affected by the calculator effect.
Anglo Saxons were like Irish and scots... That's another reason to believe they come from very similar stock. Denmark was very much like northern Britain before the Danes moved in... So people that brought in Celtic and Germanic may be identical, as both come from the same group nw of the Black Sea. Southern Britain has a good Iberian related component, pre R1b, and maybe some during the metal ages.
That would make her like 17% EEF, or there about. Are you sure that's right? See if someone's got that Dane ready to go! I'm very interested to see if he's 15% EEF, 55% WHG, and 30% ANE. Thanks, for the work David!
Never mind. I remember there were issues with the ENF. She is probably not too far from the angle/saxon. Maybe 30-35% EEF
Maybe thats an explanation for the Germans = "red haired" thing of Tacitus aswell?
"Maybe thats an explanation for the Germans = "red haired" thing of Tacitus aswell? "
I've been suspecting the samething. Northwest European-type ancestry obviously correlates with red hair. These 2 ancient samples are more extreme northwest Europeans than any today. If they are good representatives of proto-Germans, early German speakers may have had a lot of red hair.
I asked Felix of Hex 2 was a carrier of red hair, but he hasn't responded.
"I asked Felix of Hex 2 was a carrier of red hair, but he hasn't responded.:
Felix did post her blue eyes.
Some comments from the "experts" at Eupedia:
http://www.eupedia.com/forum/threads/30542-Autosomal-analysis-of-the-genomes-of-an-Iron-Age-Briton-and-an-Anglo-Saxon
They simply refuse to understand the calculator effect. It's like reading comments from the Twilight Zone.
And why are they confused by the Asian and African noise? Obviously the data's not aligned properly or there are too many duplicate SNPs.
David,
Geniuses, obviously...
Barak,
Modern English are 11% Gedrosia. Western Scots are the highest, at over 13%. As far as Europeans go.
Wouldn't that mean in fact, that this Iron Age? It pre-dates the ANglo-Saxon Invasion and settlements.
--------------------------
Telomere Length
The average telomere length from all sequence read runs gives 1.42. This means, the Hinxton-2 sample which belongs to a lady who lived 2500-1800 years back, died at the age of 65.
-------------------------
Yeah, I have no idea which period these samples are from, that's why I corrected the blog post.
If they're from the period 2500-1800 YBP then the latest ones are probably just outside the Iron Age, which I think finished around 100 AD in the UK.
We could possibly make an argument that all of these genomes are from the Iron Age, and in fact all of them might be of Celtic stock, because even the latest ones just come from the Anglo-Saxon period, rather than being verified Anglo-Saxons.
But I just don't know. To be honest, these first two look like British/Norse mixtures to me. If I didn't have anything else to go on, I'd say they were Scots or Irish with Viking ancestry from western Norway. So I can't wait to see the paper with all the archeological details.
The second one is closer to my Dad. His ancestry, is from generations of Cumbrian farmers( His dad from the North Lake District) and Scots.
Dad would score high NW too,
1 North_Sea 43.13
2 Atlantic 29.51
3 Eastern_Euro 9.14
4 West_Med 6.55
5 Baltic 5.27
6 East_Med 4.61
-- Hinxton-2 K15 --
North_Sea 45.05
Atlantic 31.08
Baltic 7.2
Eastern_Euro 12.1
West_Med 4.19
They can be from 500bce, and still be from Denmark. The dating isn't an issue, it's a question of cultural items in the burial.
We don't know anything about their burials.
All we know is that they're from 2500-1800 YBP. That does even cover the Anglo-Saxon period?
--Dads K15--
1 North_Atlantic 56.18
2 Baltic 20.79
3 West_Med 10.8
4 East_Med 6.87
5 West_Asian 2.84
-- Hinxton-2 K15 --
North_Atlantic 62.5
Baltic 25.06
West_Med 7.47
West_Asian 4.66
. Here we use ancient DNA sequencing to help address that question. We present whole genome sequences generated from five individuals that were found in archaeological excavations at the Wellcome Trust Genome Campus near Cambridge (UK), two of which are dated to around 2,000 years before present (Iron Age), and three to around 1,300 years before present (Anglo-Saxon period).
http://www.ashg.org/2014meeting/abstracts/fulltext/f140122098.htm
I know, but then we have this...
"Whole genome sequencing of 5 ancient DNA samples from skeletons excavated from Hinxton, Cambridgshire, UK with estimated dates from the iron age through the Roman period (2500-1800 years ago)."
http://sra.dnanexus.com/studies/ERP003900
"It seems like there might have been a population turnover on the Jutland peninsula after the Anglo-Saxon migrations to England,"
Bede (yes i know but) does say that. He said the Angles left the land empty. That's probably an exaggeration but interesting all the same.
It'd be awesome if some of the Hinxton samples turned out to be pure West Germanics from Jutland or Frisia from before the Danes moved in from the eastern Danish islands and southern Sweden. That'd be a real scoop.
But the most sensible assumption is that they're mixtures between native Brits and Germanics. Hence the Orcadian-like outcome. However, this obviously doesn't explain their extreme homogeneity. They don't really look admixed.
By the way, the results I'm getting with these genomes are really crisp. It's a great sign for the future. Imagine hundreds of genomes like this available online from across space and time in Europe.
David,
I saw someone say that these aren't the hinxton ring, Belgic cremation site , but from a campus in Cambridge, and covering the migration period.
Other sites too. Not just here. I have a feeling that the male is going to be Germanic.
This male has way too low of a British score, and too high in Eastern Europe, to match with modern Brits, going by MDLP k10. I haven't seen a Brit under 35% yet. His Eastern Europe is twice the British average. It appears R1b mixed with two distinct populations as far as where the hunter and farmer roots come from (Britain, only south of the Humber). On top of that, modern Norwegians have more Celto-Germanic than this individual. It appears a decent amount of British and Irish genes went back to Norway.
David,
I'm not sure if it would be of use in deciphering anything, but someone did upload the MDLP alleles for the separate tests a while back. It's post number 7 in the link.
http://www.theapricity.com/forum/showthread.php?44897-Post-your-MDLP-results
"British population history is shaped by a complex series of repeated immigration periods and associated changes in population structure. It is an open question however, to what extent each of these changes is reflected in the genetic ancestry of the current British population. Here we use ancient DNA sequencing to help address that question. We present whole genome sequences generated from five individuals that were found in archaeological excavations at the Wellcome Trust Genome Campus near Cambridge (UK), two of which are dated to around 2,000 years before present (Iron Age), and three to around 1,300 years before present (Anglo-Saxon period). Good preservation status allowed us to generate one high coverage sequence (12x) from an Iron Age individual, and four low coverage sequences (1x-4x) from the other samples."
I think that one female is more Gaul, the other Belgae, and the male is a rather unadmixed Angle, from 700CE.
It's possible that the one female is a mix of Anglo-Saxon and Brit, from 700CE. She is almost exactly like modern Southern Brits with the different components, even Iberian.
If all five turn out to be from 700CE, that still wouldn't be a surprise. As I've said before, there is a ton of evidence of native Brits holding out and being absorbed into an English identity. There are plenty of Celtic named leaders of regions in Anglo-Saxon England. I'll have to find those papers again. They list Celtic named chiefs/kings, along with archaeological evidence of British sites, next to Anglo-Saxon ones.
2 other things, that might be interesting:
1. In the "Saxon story" (10th century writer, German Saxon) there is a lil tale about where the Saxons originate. Or at least what they claim.
It says, "a long, long time ago.... in Nordland" ("Nordland" is actually the middle part of Norway), there was a famine, in the time of king Harold.
And the king said: There is not enough food for everyone, I want every family to kill all their children, but one.
But he changed his mind and said, instead of killing the children, they shall be put on boats and presented to the sea. It might bring them to better lands. And they gave the children trinkets and necklaces made of gold, that they might be able to trade.
Then the kids sailed away.... and there was a storm. The storm pushed the ships with the children against the shore of Denmark.
The older ones destroid the ships and said: Where the sea put us, we stay. We wont sail home ever.
So they wandered southward. BUt Denmark was inhabited and they kept walking south to find a place for themselfs.
In Germany they met the Thuringians.
See map:
http://upload.wikimedia.org/wikipedia/commons/6/6e/Central_Europe_5th_Century.jpg
They made a deal with the Thuringians: We want to buy terretory, for the gold we carry.
The Thuringians said: You can get as much earth, as your gold weights.... of course the story tells how the Saxons tricked the Thuringians and finaly got far more terretory than suggested.
--------------
--------------
I also stumbled over a second thing. The Germanic tribes claimed to be descants from 3 semigods. The tribes in northwest Germany claimed to descants of "Ingwio". (The tribes in Westgermany claimed to be descants of Irmin. Who is suposed to be a brother of Ingwio of course) The tribes of Eastgermany claimed "Suevo" as their father. Also one of the 3 brothers.
Now the interesting part:
There is a Scandinavian story mentioning that very semigod (with slightly different spelling. And his story is, that he got lost while on sea. And drifted far eastward.
So... if the anchestor of the northwest German tribes came by boat... from the west... huh... is the Saxon invasion of the Isles something like a backmigration? ;-)
@Fanty
I think I read - but may have misremembered - that most of the words in the Germanic languages related to boats and the sea are loan words?
Which has always made me wonder if Germanics were an amalgamation of a coastal western population and a population coming from the eastern interior who had no knowledge of boats.
Hm. Cant remember to have read something like that.
What I have read is, that Germanic is somewhat special amoung the Indoeuropean languages in Europe, for there are quiet a lot non-indoeuropean words that are also very old, so that they are possibly assimilated from an older population.
Another funny (sad?) thing is, modern German vocabulary is roughly: 30% Germanic, 60% Romance borrowed, 10% Greek borrowed. 10% other languages borrowed.
So that the number of borrowed words is huge anyways.
Lets check what an online ethymology site says about some words related to the sea:
SEA:
Old English sæ "sheet of water, sea, lake, pool," from Proto-Germanic *saiwaz (cognates: Old Saxon seo, Old Frisian se, Middle Dutch see, Swedish sjö), of unknown origin, outside connections "wholly doubtful" [Buck]. Meaning "large quantity" (of anything) is from c.1200. Meaning "dark area of the moon's surface" is attested from 1660s (see mare (n.2)).
Germanic languages also use the general Indo-European word (represented by English mere (n.)), but have no firm distinction between "sea" and "lake," either by size, by inland or open, or by salt vs. fresh. This may reflect the Baltic geography where the languages are thought to have originated. The two words are used more or less interchangeably in Germanic, and exist in opposite senses (such as Gothic saiws "lake," marei "sea;" but Dutch zee "sea," meer "lake"). Compare also Old Norse sær "sea," but Danish sø, usually "lake" but "sea" in phrases. German See is "sea" (fem.) or "lake" (masc.). The single Old English word sæ glosses Latin mare, aequor, pontus, pelagus, and marmor.
Ship:
Old English scip "ship, boat," from Proto-Germanic *skipam (cognates: Old Norse, Old Saxon, Old Frisian, Gothic skip, Danish skib, Swedish skepp, Middle Dutch scip, Dutch schip, Old High German skif, German Schiff), "Germanic noun of obscure origin" [Watkins]. Others suggest perhaps originally "tree cut out or hollowed out," and derive it from PIE root *skei- "to cut, split."
Edit:
That sums up to 110%
I think there was no "Greek" section.
30% Germanic
60% Latin AND Greek combined
10% other
I think that was it.
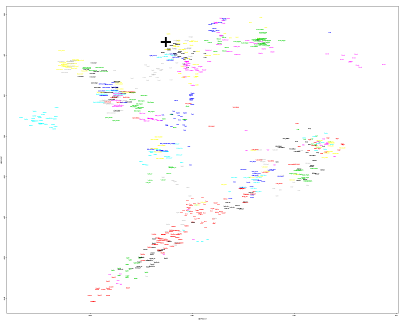
Here's a PCA of Hinxton2 ERS389796 with the Human Origins dataset. She could pass for an Icelander.
https://drive.google.com/file/d/0B9o3EYTdM8lQcWNVa051WHRacVU/view?usp=sharing
I haven't checked the f3 shared drift stats yet, but I'm betting that the Icelanders will easily top the list.
Yes, but the Icelanders are a mixture of celts and Norwegians. And the West Norwegians, are of celtic Ancestry via the Irish slaves
Did the Irish Slaves really had a big impact on modern Western Norwegians? Interestingly, the Western Norwegians have the highest North Sea score on average based on the Eutest v2 spreadsheet.
Ireland & Scotland had inferior farming land to that of South and Central England. Kind of like West Norway in climate.
So perhaps different financial gain aka slavery proper viking behaviour from the Norse. :P A few of the Modern Norwegians do carry the R-L21 Ydna. Not to sure on the age though.
It would be interesting to see how much North Sea a Doggerland individual would score. :P
@Fanty
ty, seems like i misremembered that - should have checked
I'm more interested in comparing mtdna, between Norway and the isles. I'd think female slaves siring kids is more likely than slave males breeding a lot.
"I haven't checked the f3 shared drift stats yet, but I'm betting that the Icelanders will easily top the list."
How would you explain Icelanders scoring higher WHG than Norwegians and Orcadians in Laz? Do you think it could be from extinct Hinxton-like Vikings-British or a small hunter gatherer population that lived in Iceland when Vikings discovered it? It's a crazy idea but maybe there are writings mentioning natives in Iceland like in Greenland.
Barak,
Northern Europeans may have all been Icelandic like at one time. Norwegians and such may have become more east Baltic and Saami related in more recent time.
I'd love to see the data on the birth place of immigrants to Northern Europe during the industrial revolution. Also, exact data on the amount of females taken from Finland, by Scandinavians.
It may also be quite possible that Northern Europe was only 50% Neolithic before the metal ages. Maybe a hair less..
"I'd love to see the data on the birth place of immigrants to Northern Europe during the industrial revolution. Also, exact data on the amount of females taken from Finland, by Scandinavians."
People of colonial American decent came in pre-industrial times. So, if English-Americans or Scotch-Irish in Appalachian(very isolated) show little difference to modern English and Ulster-Lowland Scots, we have to question what type of effect the industrial revolution really had.
Finnish and Sami-like ancestry would include east Asian ancestry which Swedish-Norwegian, except for northern ones, lack.
"Northern Europeans may have all been Icelandic like at one time. Norwegians and such may have become more east Baltic and Saami related in more recent time."
You mean NW Europeans or people by the North Sea coast? North Sea people today aren't just more eastern than Hinxton 1 and 2, but also more southwestern. We only have 2 early medieval individuals from England, they don't represent everyone during that time or shortly before from Ireland-Sweden. We might find more surprises in the future.
I'm sure French are close to no different from the Gauls Caesar conquered. Iberians also have probably had little foreign admixture since that time. Central Europe and Britain is defiantly what have changed the most.
David,
Any confirmation yet on which samples these are? Are they the Cambridge or Hinxton remains?
CEU, in many runs, does show up in the British plot. These people are mostly of British and North German descent, I believe.
It is very possible that Danes made Denmark closer to modern Scandinavians, instead of Orcadians and Icelanders. I show up as SE English, 100% English in Dr McDonald's run. My ancestry is something like just over 1/2 English, Scot, Welsh, and Irish, 1/8 Danish, 1/8 Pomeranian German, 5/64 Alsatian, 1/16 Bavarian, The rest is Jew and various other parts of Western and Central Europe. I show minor Iberian and Mediterranean in most test companies.
We probably won't know for sure who these samples represent until the paper gets published. But anyway, here are the updated oracle results for Hinxton1.
https://drive.google.com/file/d/0B9o3EYTdM8lQdDYzMEF6Zi1qUWc/view?usp=sharing
https://drive.google.com/file/d/0B9o3EYTdM8lQRERCMDgtcks1OXM/view?usp=sharing
And here are the results for Hinxton2.
https://drive.google.com/file/d/0B9o3EYTdM8lQQVRfWXdhQXVpVE0/view?usp=sharing
https://drive.google.com/file/d/0B9o3EYTdM8lQSnpRcmxqWURzSmM/view?usp=sharing
Where's Icelandic on your K13 and K15 spreadsheets?
I just asked Felix. He said that it is from the Iron Age group of Belgae. This still leaves room for natives living under Belgae rule to be much more EEF, like modern English people. Unless Felix is mistaken, but I guess we will wait until the paper comes out.
It's there now. Also, here are some individual results.
https://docs.google.com/spreadsheets/d/1bSB8iAVjEynMnGdcpoowuZTUF5mY3Sl0EfYTS8ElUgo/edit?usp=sharing
https://docs.google.com/spreadsheets/d/1P1sd-LthAze4rdYH--uHMtE8CC8ypdS_wzhc0KGY120/edit?usp=sharing
I spoke with Felix, again. I showed him the Cambridge abstract, and he is now not sure where the samples are from. Looks like we're waiting for a paper. Any idea when it's supposed to be available? Is it part of the conference coming up?
I went ahead and e-mailed Dr. Schiffels. He is out of the office until the 16th. Hopefully, I get a reply back in a few days.
David,
Felix has Gok4 and Gok5. You should run those and see if either is pure enough to make an EEF cluster with Stuttgart, for the k6.
Well, someone told me that these are the Anglo-Saxons
Hinxton 1 - ERS389795 - M - R1b-L11 - K1a1b1b - Anglo-Saxon Low 1-4x
Hinxton 2 - ERS389796 - F - - H2a2b1 - Anglo-Saxon low 1-4x
Hinxton 5 - ERS389799 - F - - H2a2a1 - Anglo-Saxon low 1-4x
So, 3 and 4 are Brits.
2 and 5 could be hybrid Brit and Anglo-Saxon. Their Iberian is half of 3 and 4, on MDLP. 1 looks pretty pure, and I'd bet an Angle.
Hinxton2 is clearly more northern than Hinxton1. Nothing Iberian about it at all.
They score the Iberian component, where as it's not in 1, and 2 and 5, are half of 3 and 4. The paper didn't say they were like Iberians but had more Iberian relation than the Saxons, and that all samples fit as Northern European.
Are you talking about the results posted by Genetiker?
His results look very noisy, which is strange, because the two Hinxton files I have show hardly any noise.
I don't think the ancestry proportions posted by him are accurate.
It does match though... Celts more Iberian, Saxons more balto Finnic. 2 Brits, 2 1/2 Brit, one full anglo saxon.
There's no way that Hinxton2 can be more Iberian than Hinxton1.
Hinxton 2 can be more Iberian related and have 2 percent less EEF than the anglo-saxon. All it takes is more mesolithic survival in ancient Britain than in Denmark. Wait until Hinxton 3 and 4, to make any calls yet.
Hinxton 2 looks 42% EEF, 41% WHG, 17% ANE. Hardly any difference.
If hinxton 3 and 4 are further from Britain, you'll have your answer. Only the Saxons were close to modern Brits.
If hinxton 2 is a hybrid, as it looks, it should be further from Brits than hinxton 1. Do you have the files from hinxton 3 yet?
"If hinxton 3 and 4 are further from Britain, you'll have your answer. Only the Saxons were close to modern Brits."
Since the abstract says the Anglo-Saxon samples were c loser to modern Brits(including English), Hinx 1 and 2 should be Iron age Brits.
Barak,
You can't say that without seeing 3 and 4. Native Brits could've been Icelandic like. L11 makes it unlikely. Plus, word is that 3 or, I forget which, is L21 or DF21.
Post a Comment