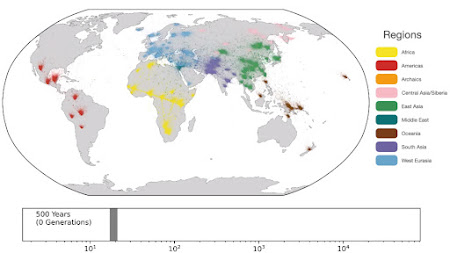
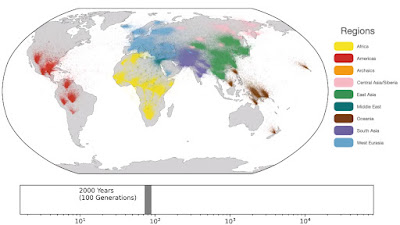
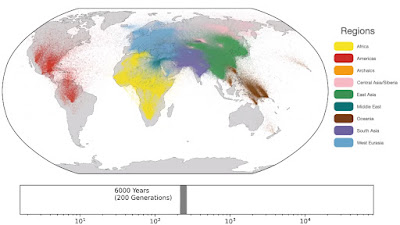
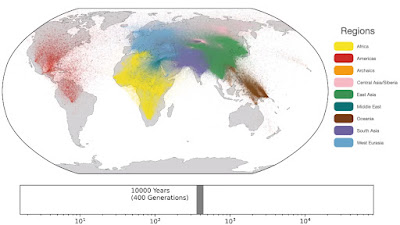
Spatio-temporal dynamics in human history. This movie shows the estimated geographic locations of ancestors of Human Genome Diversity Project, Simons Genome Diversity Project, Neanderthal, Denisovan, and Afanasievo samples over time. Each dot represents an edge in the tree sequence of chromosome 20, where the time and geographic location of the parent and child nodes of the edge have been estimated. The locations of edges at each point in time are plotted along the great circle between the parent and child nodes. Edges are colored by the region of the descendants of the child node. If an ancestral lineage has ancestors in multiple regions, its color is the average of the respective colors of each region.See also... Haplotype-based PCA of West Eurasia and Europe
Thursday, February 24, 2022
A unified genealogy of modern and ancient genomes (Wilder Wohns et al. 2022)
Over at Science at this LINK. Broadly speaking, this looks like a more sophisticated version of something that I tried about five years ago (see here).
I wonder if they got the idea from me? Honestly, I wouldn't be surprised if they did. But like I say, their methods are way more advanced.
Keep in mind, however, that for now, their analysis includes 3601 modern genomes and just eight ancient genomes. That's because they can only run super high quality ancient sequences. The ratio of ancient genomes will no doubt rise rapidly over the next few years, and that's when things will get really interesting.
Below are some screen caps from a clip accompanying the paper, freely available here. This is the caption to the movie:
36 comments:
Read the rules before posting.
Comments by people with the nick "Unknown" are no longer allowed.
See also...
New rules for comments
Banned commentators list

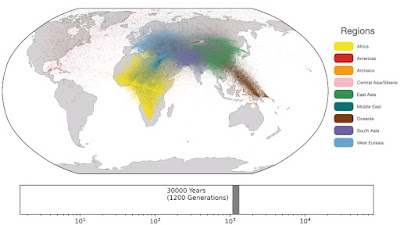
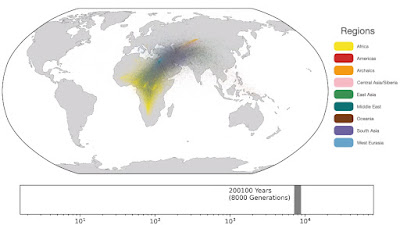
Interesting paper but what are the graphs in the end showing? Native Americans sharing ancestry with Eurasians only 2000 years ago and Eurasian ancestry dating to Africa only 10000 years ago? Are these not under-estimates or am I misreading this?
ReplyDeleteAlso how can Papuans owe so much of their ancestry to East Asians only 2000 years ago already?
Not really sure what to make of all the dots on the oceans in the earlier diagrams. What does that mean?
ReplyDeleteThey're not literally in the ocean. They're placed there as a consequence of trying to match genes with geography within such a broad context.
ReplyDeleteIn part, that's why I said adding many more ancient sequences will be interesting, because then it'll be possible to work out precise geographic links and migration routes.
I don’t think this represent an actual genealogy of Eurasian populations, just the programme’s rationalization of genetic diversity. Although some sub branches might be genuine affinities
ReplyDeleteI suspect some might be misled if they take the results at face value
@Rob
ReplyDeleteDid you notice in their supplementary figures how African pops seem to be more Denisovan than South Asians or how a subset of Japanese are more Afanasievo than northern Han?
I didn't expect Japanese to even have Afanasievo admixture though some may have trickled in. Still how can it be more than northern Han...
New paper from the Reich team in Nature (free)
ReplyDelete"Ancient DNA and deep population structure in sub-Saharan African foragers"
https://www.nature.com/articles/s41586-022-04430-9
Since it’s kinda quiet, can I ask a dumb layman question about the tree diagram? (Answers that would suit a ten-year-old, please).
ReplyDeleteRathlin 1 & 2 (Rathlin being the stepping-stone island between Scotland and N Ireland) — we were told that they were close kin, no more than one or two generations apart. The dating (c. 2000 BCE) almost completely overlaps, allowing them to potentially be exact contemporaries. These were apparently among the earliest generations of “Rhenish” beakers to the British Isles. You will know that a fair percentage of Irish and Scottish men come from the same sub-clade of L21 they did (DF21 & its own descendant lineages).
We (the public) were being told a few years ago that the mutations that define DF21 happened pretty close to the time (whether before or after) that this Beaker group left the continent.
The branches on the tree diagram where Rathlin 1&2 are represented are close but distinct — neighbours labelled Afanasievo & Yamnaya/Kalmykia. (I’m even more confused about the RISE labels; I had thought that meant burials around Rostov on Don, but….?)
Can you help me understand why each Rathlin guy is on a distinct branch of this diagram? It makes sense that these two would be very close together, but why are they not just together on the same branch?
The paper is behind a paywall but they probably relied on short IBD segments and varied the lengths of the shared segments to come with a time scale of shared drift. We have done a ton of research on this over the past few years and I still have one of our robust servers crunching the numbers 24/7.
ReplyDeleteThe problem is far greater than most researchers realize. Most researchers will naively run with a dataset not realizing the effects of the sequence read aligner and other components used calling genotypes and even bigger, the effect on changing the ascertained SNPs on the downstream results whether they’re using PCA, formal stats, IBS or IBD.
Based on my experience processing DNA reads and downstream analysis for the past few years I have found that changing any of the following, changes sometimes drastically the IBD, IBS, formal stat and PCA results:
1- The DNA sequence pipeling and variant calling method. This is especially true with aDNA because the reads are shorter and tend to map to wrong chromosomes more often and also because of deamination. Lately we have been using the older slower bwa aln and disabling seeding with -l1024 and setting maximum edit to -n0.01 and maximum number of gap opens to -o2 to reduce Hg37 reference bias. Martiniano et al recently published something relating to this "Removing reference bias and improving indel calling in ancient DNA data analysis by mapping to a sequence variation graph".
2- Ascertainment bias we have accumulated a ton of analysis results, just not had the time to publish, where we performed IBS and IBD using the Affymetrix Axiom 1240K set vs WGS vs Illumina Omni2.5 2.3M (1000G) vs Illumina Omni 5.4 4.3M (HapMap). Results significantly varied whether we did PCA, IBS or IBD.
You'll routinely read that humans have 1.5M or 2M polymorphisms which sort of corresponds to the amount of SNPs when you do WGS on someone. Although this may be true for any certain individual, the polymorphic sites vary from individual to individual, and population to population such that if we total up all the polymorphic sites we have discovered for ALL people we have genotyped to date they may easily exceed 350 million sites.
The big problem which I don't think will be solved anytime soon is which of the 350 million intersecting polymorphic sites do we select when performing analysis. No matter which ones we choose there will always be some bias. Thus even WGS analysis is biased.
I think it will be a long time before most reasearchers become cognizant of everyting I just wrote and with time you’ll see relationships between populations change
I must confess I don't really understand what they're doing here. Nonetheless, some interesting points from the pre-print:
ReplyDelete1: "Traversing towards the present, by 280 kya, the centre of gravity of ancestors is still located in Africa, but many ancestors are observed in the Middle East and Central Asia and a few are located in Papua New Guinea. At 140 kya, more ancestors are found in Papua New Guinea. This is almost 100 kya before the earliest documented human habitation of the region. However, our findings are potentially consistent with the proposed timescales of deeply diverged Denisovan lineages unique to Papuans. At 56 kya, some ancestral lineages are observed in the Americas, much earlier than the estimated migration times to the Americas. This effect is likely attributable to the presence of ancestors who predate the migration and did not live in the Americas, but whose descendants now exist solely in this region; the same effect may also explain the ancient ancestors within Papua New Guinea. Additional ancient samples and more sophisticated inference approaches are required to distinguish between these hypotheses. "
2: "(O)ur approach requires phased genomes, which is a particular challenge for ancient samples that typically pick random reads to create a “pseudo-haploid genome”.
However, it should be possible to use a diploid version of the matching algorithm in tsinfer to jointly solve phasing and imputation. This also has the potential to alleviate biases introduced by using modern and genetically distant reference panels for ancient samples. Recent work focusing on inferring genealogies for high-coverage ancient samples, and using mutations dated in such a genealogy to infer relationships of lower coverage samples through time, offers an alternative strategy for accommodating the unique challenges of ancient DNA in this context.
In addition, our approach to age inference within tsdate only provides an approximate solution to the cycles that are inherent in genealogical histories and there are many possible approaches for improving the sophistication of spatio-temporal ancestor inference.
Re; what they have done with the Afanasievo samples, since there are starting to be quite a few family groups found now, I wonder if there is any potential to enrich samples from e.g. Hazleton Long Cairn for phasing*, or if this would even have any scientific value?
ReplyDelete----
*Another upcoming example: https://journals.openedition.org/bmsap/9242 -
"Exploring Neolithic social structures using genetic structure of two large families at Gurgy “les Noisats”, France" - "We obtained genomic data for 94 of the 128 individuals from the site. We have reconstructed two large genealogies, one of which covers 7 generations and brings together 63 individuals.")
(Off topic but some more upcoming things from France noted here: https://rmpr.fr/wp-content//uploads/sites/7/2022/01/RNS4_LaRochelle_PreProgramme.pdf-
"Between continuity and discontinuity - Contributions of genomics to understanding biological and cultural dynamics of Neolithic farming communities from Languedoc" - A. Arzelier, M. Rivollat, H. De Belvalet, M.-H. Pemonge, D. Binder, F. Convertini, H. Duday, M. Gandelin, J. Guilaine, W. Haak, M.-F. Deguilloux, M. Pruvost
"Paleogenomic analysis of a collective burial from the Final Neolithic of Paris Basin sheds light on population processes during the 3rd millennium BC" - O. Parasayan, C. Laurelut, A. Corona, C. Domenech-Jaulneau, T. Grange, E.-M. Geigl
and https://www.sapweb.fr/images/Programme_SAP2022.pdf
"Biological interactions between hunter-gatherers and Neolithic farmers: the remarkable case of Southern France communities" - Ana Arzelier, Maïté Rivollat, Harmony De Belvalet, Marie-Hélène Pemonge, Didier Binder, Jean Guilaine, Fabien Convertini, Muriel Gandelin, Henri Duday, Wolfgang Haak, Marie-France Deguilloux, Mélanie Pruvost - In order to overcome some of these gaps, we sequenced the genomes of 31 ancient individuals dated between 5500 and 1000 BC from six sites in the Occitanie region illustrating various archaeological contexts and funerary practices. ... We thus detect high proportions of Mesolithic ancestry in the Early Neolithic groups of southern France, in contrast to neighboring regions of Western Europe. We highlight contrasting scenarios between the different currents of Neolithization in terms of migratory processes and intergroup interactions. These results also highlight the persistence of significant hunter-gatherer ancestry in several Western European groups during the Final Neolithic, highlighting the complexity of regional demographic processes.
New radiocarbon and paleogenetic data on the Villard dolmen (Lauzet-Ubaye, Alpes-de-Haute-Provence) - Aurore Schmitt, Fabien Convertini)
So did the original Homo Sapiens go extinct ? Mmm...Looking at all the deep diverging ghost population admixture in all, it surely seems so...
ReplyDelete@Wee e
ReplyDeleteThat tree I posted is based on IBD segments over a certain length.
So it seems that what's happening there is that one of the Rathlin samples shares more of these large segments with one of the Afanasievo samples than with the other Rathlin sample. That is, they appear to have a close genealogical relationship.
If I was to reduce the size of the segments, and indeed, if I was to base the tree on unlinked genotypes, then the two Rathlin samples would form a branch to the exclusion of the Afanasievo sample, which would cluster with the Yamnaya samples.
A potential confounding factor here is that the genotypes in the ancient genomes that I used were heavily imputed.
So there might be a close genealogical relationship between the Rathlin sample and the Afanasievo sample, but then again, this may well be a mistake because of the low quality of the data used.
Also a good article
ReplyDelete“ Neolithization and Population Replacement in Britain: An Alternative View”
Highlights the problems of the Brace/ Reich narrative
You mean the alternative view that 90% + of the ancestry in Britain changed due to gradual drift?
ReplyDeleteAnd of course there really was no migration, because migrations didn't happen until the 20th century.
Drift is not the premise. The issue is the autosomal reductionist framework through which some Labs operate.
ReplyDeleteno, British farmers aren't from the Near East. To say so is to misunderstand, at best, or falsify history at worst. As Thomas points out, this is an act of Atlantic foragers 'going Neolithic', riding the Neolithic boom, and acquiring EEF ancestry from Paris basin Farmers.
This reality is very different to the one Brace et al spells out in their effort
gamerz_J,
ReplyDeleteR-Z2103 (TMRCA 4910 years) currently has a total of 116 members on 23mofang.
Their geographic distribution is as follows:
Shandong x13
Xinjiang x12
Shanxi x11
Beijing x7
Hebei x7
Anhui x6
Gansu x5
Jiangsu x5
Liaoning x5
Fujian x4
Heilongjiang x4
Inner Mongolia x4
Shaanxi x4
Sichuan x3
Guangdong x2
Henan x2
Hubei x2
Jilin x2
Ningxia x2
Tianjin x2
Chongqing x1
Jiangxi x1
Taiwan x1
Italy x1
Turkey x1
Ukraine x1
USA x1
undisclosed x7
So, while possible patrilineal descendants of males of the Afanasievo culture do appear to be relatively common in Xinjiang as one might expect, they may be found sporadically in almost any part of China. They also seem to be relatively common in the northeastern part of China proper: Shanxi, Shandong, Hebei, Beijing, Tianjin.
Overall, however, R-Z2103 is rare in China (≈0.05% or about one in every two thousand Chinese males).
Among subclades of Y-DNA haplogroup R1b, R-PH155 (≈0.08%), R-M478 (≈0.07%), and even R-L51 (≈0.11%) are more common than R-Z2103 at 23mofang. However, most members of R-L51 at 23mofang appear to be actual foreigners or descended from foreigners (e.g. English, Portuguese) within the last several generations, so their historical connection with China appears to be nonexistent or extremely shallow.
@Rob
ReplyDeleteWhen these British archeologists say drift, they don't mean genetic drift, but a gradual genetic shift over time.
So they're really talking about isolation by distance.
The problem with applying this concept to Late Neolithic Britain is that the appearance of Bell Beakers was actually because of a migration, and not some gradual change in genes and culture due to influences from nearby parts of Europe.
I saw the paper existed but didn't read it. I don't think the author talks about LNCA genomic replacement at all.
ReplyDeleteDave ; they’re taking here about the early Neolithic; not the “drift hypothesis” of British beakers.
ReplyDeleteWe agree that the latter was a relatively sudden; truly sweeping event
OK, we're talking about different things.
ReplyDeleteThe "Steppe Drift Hypothesis" is really just saying "Were there more migrants with steppe-ancestry after 3000 BCE and before 2600-2500 BCE, and we can't see them because they adopted local cremation?" and thus whether cultural continuity of some practices might be higher than we'd think from a sudden replacement that happens only when Beaker burials emerge.
ReplyDeleteI think it's halfway plausible, but I don't know and there are good reasons to think its doubtful; single burial inhumation seems very culturally important to the CWC/Beaker group, so I would have thought that surely some migrants would have insisted on retaining it, and have cultural resistance to assimilating with people who did not practice it (perhaps more resistance than say, with people in SE Europe who already practiced simialr forms of inhumation?). Like, maybe there were these steppe offshoots who showed up, were awed by the network of people across Southern Britain who were building megalithic structures still (or worshipping at them at least) and practicing cremation, and they joined these groups and largely replaced much of the genomic ancestry, then to merge with later truly Beaker migrants... But I don't think we have any evidence for this, and the strong adherence to the single-grave burial has suggestion against it.
@Davidski
ReplyDeleteThanks for replying. I had a look at Cassidy’s paper (which I am not equipped to understand). Rathlin 1 coverage is 10-11x but Rathlin 2 is about 1.5x.
@ Matt
ReplyDeleteOnly thing still left to answer though is if CWC/ EKG arrived in Rhine as early as 2800 bc, why did it take them 400 years to reach Britain (a/p the more commonly accepted 2400 bc date). Crossing the Atlantic was a difficult task ? but had obviously been done before by Mesolithics & Neolithics
@Dilawer
ReplyDeleteBased on your own research, do you think that there is some fundamental misunderstanding of human population genetics/history, given these apparently unaccounted for complications you seem to have found? What are some novel insights you think you've found that the mainstream has missed?
@ Rob
ReplyDelete"Only thing still left to answer though is if CWC/ EKG arrived in Rhine as early as 2800 bc, why did it take them 400 years to reach Britain (a/p the more commonly accepted 2400 bc date). Crossing the Atlantic was a difficult task ? but had obviously been done before by Mesolithics & Neolithics"
The climate at the time might have been a factor. Although the following time period slices are very broad brush stroke, it could indicate what aspects of climate change either attracted, or pushed out, people migration across bodies of water, and whether water levels facilitated island hopping, perhaps allowing people with less developed water craft and skills to get across during favorable periods.
5000 - 3000 BC; Climatic optimum; Warm conditions; temperatures were perhaps 1 to 2 degrees Celsius warmer than they are today. Great ancient civilizations began and flourished.
3000 - 2000 BC; Cooling trend; drops in sea level and the emergence of many islands.
2000 - 1500 BC; Short warming trend.
1500 - 750 BC; Colder temperatures and renewed ice growth, sea level drop of between 2 to 3 meters below present day levels.
http://www.atmo.arizona.edu/students/courselinks/fall12/atmo336/lectures/sec5/holocene.html#:~:text=By%205000%20to%203000%20BC,Optimum%20or%20the%20Holocene%20Optimum.
@Rob, yes, that seems true. I don't really have any ideas about it though, you?
ReplyDelete@Robb.
ReplyDeleteBuilding speculatively on to MAD’s answer, it could have taken centuries for land-hunger to build up once the steppe descendants reached the continental west coast. Then there’s the actual going to and thriving within the islands.
The logistics and the risk/benefit calculation for a herder or farmer community migrating as a community across the Channel is much tougher than the same community migrating on the Continent. Expensive, takes total commitment, and no bridge to burn. Herders need agrarian farmers to trade with. Agrarian farmers also need a wider infrastructure: and a social support structure for the “two years in ten” when crops fail or animals sicken. The prospect of being cut off from both (in a famously unpredictable climate) might not be enticing, whether that’s in an “empty” island or one to be wrested from Anatolian-descended farmers. You’d need a push as much as a pull. They would hear about our fatally unpredictable weather.
The Viking Greenland colonies were still few and small after centuries, even given a climatic rosy period: when they realised that the next ship could be the last to bother coming, a few last Greenland-born arrived hungry and exhausted in Iceland. And no-one heard from the few holdouts again. (Until now, when remains have been found still in their homes, even their beds.) They certainly could have survived if they had been willing to adopt the “skraeling” way of life, and they had centuries to try.
But Britain and Ireland weren’t quite that harsh, as there were more options if trade fell off or the climate hiccupped. Especially for people with the lactase persistence gene.
An industrial/trader outpost here and there trading for a few calves and a couple of adventurous herdboys here, advertising for a good brewer or potter spouse there — slow growth, satellites either upriver or shipped around the coast. Keeping up connections, they may have had more of an “expat” than insular identity for centuries. The norm may not have been emigration but a few years of manly adventure, a fortune to be made (I’m thinking of British small-merchant sons in Demerara & Essequibo here, attracted by swashbuckling tales of the West Indies and actually living as accountants/managers on isolated plantations, terrified of slaves and dying in their droves of boredom, alcoholism and fever. But maybe Beaker lads were thinking of roving wolf-cub bands, even if their actual job was digging copper ore.) It might have been boring and high risk - and the ambition might have been to get rich quick, then back home to buy a cattle herd.
The later population explosion could have originated in comparatively few people that maybe kinda got stuck here despite themselves, or whose kids decided to stay when the parents went back to the “old country”.
It has puzzled me why in these islands we have quite a piecemeal, heterogenous variety of Beaker remains, (different from one valley to the next, here in Scotland) yet we have this insanely limited set of haplotypes (male and female) from them. Maybe most Beaker settlements here failed. Maybe for the same reasons the predecessor (Anatolian-descended) population dwindled; whatever reason that was.
I’m not arguing that any if this happened: just that there are tons of scenarios, not nevessarily mutually exclusive, to be considered.
@Rob
ReplyDeleteRemember, in the mesolithic, Britain was a peninsula, not an island. Right up to about 6,500BCE.
Even after that, what is now Dogger Bank was still an island above water — a pretty extensive stepping stone refuge / navigation aid &/or overnight rest — until about 5,000BCE.
All agreeable possibilities.
ReplyDeleteIt just seems that 'Beaker people' started expanding as a concerted phenomenon ~ 2500 bc, soon after including Britain, although their predecessors had arrived to Europe already by 2800 bc
The main climactic event in M3 was ~ 2200 bc.
@ Wee
Yep for British farmers, they did have a smoothing out of growth c 3000 bc, and a return to hunting. Some have described it as a bust, but IMo it's more likely that steady state of growth was reached & diversification of economy perhaps. The entire Island shared had shared similar ceramic styles and construction of Stonehenge, and so forth as the earliest Beaker people arrived. So they don't immediately seem as a people in crisis, quite the contrary, but then again creating large offerings could have been an act of appeasement and asking for favour from the Gods ?
British Beaker was variable in the burials, but its all pulled out of a common repertoire.
@Ebizur
ReplyDeleteThanks for the rundown on the haplogroups, I was mostly pointing out that it seems odd for Japanese to have more Afanasievo-related ancestry than northern Han based on broad West Eurasian affinities found in northern Han in previous papers, just like it is odd that African pops evince more sharing with Denisovans compared to South Asians (that really should not be the case) unless in both cases they are mis-infering shared variants due to old common descent as admixture.
@Dilawer
They are using tsinfer, not IBD. In my understanding it's quite a robust software/program.
Dilawer, does this mean that all the results of archaeogenomic research to date are unreliable?
ReplyDeleteDavidski, whats your best admixture model for Latvia_BA?
ReplyDelete@ambron
ReplyDeleteDilawer, does this mean that all the results of archaeogenomic research to date are unreliable?
Absolutely not.
And if you believe so, then there's no hope for you.
@David
ReplyDeleteAre Dilawer’s grievances legitimate? I don’t know how to analyze genomes directly with computer programs, so I’m not sure how to interpret anything he said.
@Simon
ReplyDeleteHe's exaggerating the problems.
One obvious way to check that even low quality ancient genomes are a reliable source of data is to compare them to modern genomes that plausibly they should closely resemble based on their geographic origins and archeological contexts.
And, of course, when we do that then everything always makes sense. I've proved this on countless occasions at this blog.
For instance, ancient Germanic and Celtic samples look exactly like we expect them to look, and so on and so forth.
No one in their right mind will claim that this is a coincidence, or that we should expect different outcomes...
https://eurogenes.blogspot.com/2019/12/avalon-vs-valhalla-revisited.html
The only people who do make a fuss about this to the point of claiming that archaeogenetics is not a reliable science are those who don't like the results that they're seeing for one reason or another.
What's most striking to me in the video clip visualizing the data is that per its color code it distinguishes between ancestors of West Eurasians (depicted as in the Levant) and ancestors of Subsaharan Africans already by before 300,000 years ago. This would seem to imply that proto-West Eurasians and proto-Black Africans were already separated, divergent lineages even before the commonly accepted timeline of the original speciation of sapiens, and would also necessarily refute the mainstream narrative of a recent Out of Africa scenario.
ReplyDelete