search this blog
Wednesday, May 18, 2022
Geography is hard (for some)
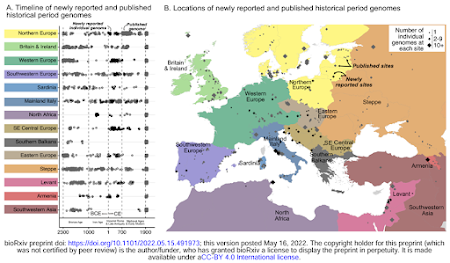
It's that time of the academic year again when bioRxiv is inundated with ancient DNA preprints. I'm not complaining, but I almost spat out my coffee when I saw this map in one of the new manuscripts (here).
What's the logic behind labeling almost all of Eastern Europe as "Steppe", and instead labeling just Czechia, Hungary and Slovakia as "Eastern Europe"? In my opinion those three countries, plus Poland, are better described as East Central Europe anyway.
It seems to me that many people working at the highest level in population genetics simply don't know what the Eurasian steppe is. They appear to see it as a continent of its own, when, in fact, it's a topographical feature and ecoregion that straddles the continents of Europe and Asia. That's why it's called the Eurasian steppe, and it's made up of three main parts: the Pontic-Caspian steppe of Eastern Europe, the Kazkah steppe of Central Asia, and the Eastern steppe of Mongolia.
Here's the same map with a few corrections (in red). Much better, don't you think?
Citation...
Antonio et al., Stable population structure in Europe since the Iron Age, despite high mobility, bioRxiv, posted May 16, 2022, doi: https://doi.org/10.1101/2022.05.15.491973
See also...
Matters of geography
Tuesday, May 17, 2022
Genome-wide data from medieval German Jews (Waldman et al. 2022 preprint)
Over at bioRxiv at this LINK. Here's the abstract:
We report genome-wide data for 33 Ashkenazi Jews (AJ), dated to the 14th century, following a salvage excavation at the medieval Jewish cemetery of Erfurt, Germany. The Erfurt individuals are genetically similar to modern AJ and have substantial Southern European ancestry, but they show more variability in Eastern European-related ancestry than modern AJ. A third of the Erfurt individuals carried the same nearly-AJ-specific mitochondrial haplogroup and eight carried pathogenic variants known to affect AJ today. These observations, together with high levels of runs of homozygosity, suggest that the Erfurt community had already experienced the major reduction in size that affected modern AJ. However, the Erfurt bottleneck was more severe, implying substructure in medieval AJ. Together, our results suggest that the AJ founder event and the acquisition of the main sources of ancestry pre-dated the 14th century and highlight late medieval genetic heterogeneity no longer present in modern AJ.It's nice to finally see some ancient Jewish genotypes on the way, but there's a bit of a problem with this preprint. The fact that the authors are using modern-day Russians to model Eastern European-related ancestry in these Ashkenazi ancients from Central Europe tells me that they're somewhat confused. They did this because some of the Jews harbor significant Slavic ancestry and minor but perceptible East Asian ancestry, and Russians are Slavs who carry some Siberian ancestry, which is closely related to East Asian ancestry. Thus, broadly speaking, in terms of the right mix of DNA, Russians do the job. However, as per the preprint, based on historical data, these Jews probably sourced their Slavic ancestry from Bohemia, Moravia and/or Silesia, and the Slavic speakers in these regions carry very little, if any, East Asian or Siberian ancestry. I'm sure the authors can verify this claim without too much trouble. Ergo, it's likely that the Erfurt Jews received their Slavic and East Asian admixtures from different sources, and possibly at different times. I'd like to see Waldman et al. tackle this issue properly. I suspect that if they do, they might discover something interesting and perhaps unexpected about the ethnogenesis of Ashkenazi Jews. See also... My take on the Erfurt Jews Mediterranean PCA update
Saturday, May 7, 2022
Population genomics of Stone Age Eurasia (Allentoft et al. 2022 preprint)
Over at bioRxiv at this LINK. It'll take me a few days to read this manuscript properly. Here's the abstract:
The transitions from foraging to farming and later to pastoralism in Stone Age Eurasia (c. 11-3 thousand years before present, BP) represent some of the most dramatic lifestyle changes in human evolution. We sequenced 317 genomes of primarily Mesolithic and Neolithic individuals from across Eurasia combined with radiocarbon dates, stable isotope data, and pollen records. Genome imputation and co-analysis with previously published shotgun sequencing data resulted in >1600 complete ancient genome sequences offering fine-grained resolution into the Stone Age populations. We observe that: 1) Hunter-gatherer groups were more genetically diverse than previously known, and deeply divergent between western and eastern Eurasia. 2) We identify hitherto genetically undescribed hunter-gatherers from the Middle Don region that contributed ancestry to the later Yamnaya steppe pastoralists; 3) The genetic impact of the Neolithic transition was highly distinct, east and west of a boundary zone extending from the Black Sea to the Baltic. Large-scale shifts in genetic ancestry occurred to the west of this "Great Divide", including an almost complete replacement of hunter-gatherers in Denmark, while no substantial ancestry shifts took place during the same period to the east. This difference is also reflected in genetic relatedness within the populations, decreasing substantially in the west but not in the east where it remained high until c. 4,000 BP; 4) The second major genetic transformation around 5,000 BP happened at a much faster pace with Steppe-related ancestry reaching most parts of Europe within 1,000-years. Local Neolithic farmers admixed with incoming pastoralists in eastern, western, and southern Europe whereas Scandinavia experienced another near-complete population replacement. Similar dramatic turnover-patterns are evident in western Siberia; 5) Extensive regional differences in the ancestry components involved in these early events remain visible to this day, even within countries. Neolithic farmer ancestry is highest in southern and eastern England while Steppe-related ancestry is highest in the Celtic populations of Scotland, Wales, and Cornwall (this research has been conducted using the UK Biobank resource); 6) Shifts in diet, lifestyle and environment introduced new selection pressures involving at least 21 genomic regions. Most such variants were not universally selected across populations but were only advantageous in particular ancestral backgrounds. Contrary to previous claims, we find that selection on the FADS regions, associated with fatty acid metabolism, began before the Neolithisation of Europe. Similarly, the lactase persistence allele started increasing in frequency before the expansion of Steppe-related groups into Europe and has continued to increase up to the present. Along the genetic cline separating Mesolithic hunter-gatherers from Neolithic farmers, we find significant correlations with trait associations related to skin disorders, diet and lifestyle and mental health status, suggesting marked phenotypic differences between these groups with very different lifestyles. This work provides new insights into major transformations in recent human evolution, elucidating the complex interplay between selection and admixture that shaped patterns of genetic variation in modern populations.Allentoft et al., Population Genomics of Stone Age Eurasia, bioRxiv, posted May 06, 2022, doi: https://doi.org/10.1101/2022.05.04.490594 See also... Understanding the Eneolithic steppe
Subscribe to:
Posts (Atom)