search this blog
Saturday, May 28, 2016
Indian genetic history in three simple graphs
Enjoy, but please, those of you still sore about the passing of the Out of India, Out of Armenia, Out of Your Hat, or indeed, Out of Your Ass Indo-European hypotheses, try not to fill up the comments with the usual inane drivel. Thanks in advance for your cooperation.
Tuesday, May 24, 2016
Corded Ware women more mobile than their men (Sjögren et al. 2016)
From a new paper at PLoS ONE on the diet and mobility of Corded Ware (CW) people in Central Europe (emphasis is mine):
Regarding the formation of the CW, some archaeologists point out the contribution of different regions to the material set of the "CW-network", while others note similarities with the steppe, in particular with the Yamnaya culture, as a possible area of origin. This is based on similarities in burial rituals. Some authors have suggested that this culture practiced a form of mobile pastoralism, which spread towards the west through migration and/or cultural influence, and gave rise to the CW. In the process, Indo-European language would also have spread over Europe [3, 4, 5, 6, 7]. Recently, these hypotheses have gained support from aDNA studies of Yamnaya and CW burials. Allentoft et al [8] and Haak et al. [9] show that a genetic transformation took place in areas where previous Neolithic DNA was heavily reduced and complemented by Yamnaya DNA. This new genetic presence was lasting and provided much of the genetic material for contemporary European populations. There is increasing evidence for some kind of population reduction or crisis toward the end of the middle Neolithic facilitating this introduction of new genes (e.g. [10]) and recent research has documented the presence of plague among Yamnaya and Corded Ware individuals [11], which may have spread among Neolithic populations prior to the migrations. ... The number and proportion of females with distinctive strontium isotope ratios is notable and suggests that women were more mobile than males in CW society. Such evidence fits well with recent genetic information documenting more varied haplogroups among CW females [14]. Müller et al. [2] suggest female exogamy as a means of maintaining lineage identify in the face of rapid, long-distance mobility. Haak et al. [25] also reported genetic and Sr isotope ratio differences between males and females at Eulau, Germany, suggesting female exogamy. The fact that such a difference is identifiable at all also suggests that males were largely stationary, at least in the sense that they were mostly born, raised and buried in the same locality. We suggest that this reflects a stable exogamic system where women moved to their husband’s settlements, existing at Bergrheinfeld for several generations. As no distinctions in burial treatment were associated with incoming women, either the exogamic exchange involved only CW groups, or incoming women were completely integrated into CW society.Citation... Sjögren K-G, Price TD, Kristiansen K (2016) Diet and Mobility in the Corded Ware of Central Europe. PLoS ONE 11(5): e0155083. doi:10.1371/journal.pone.0155083 See also... Female mobility and exogamy as the main drivers of foreign admixture during the Late Neolithic/Early Bronze Age shift in Central Europe (Knipper et al. 2017)
Sunday, May 22, 2016
Parallel migrations in brown bears and humans within Eurasia
If, as per the abstract below, a major part of the Native American gene pool derives from the Altai-Sayan refugium, then it's likely to be the Ancient North Eurasian (ANE) part, rather than the East Asian part. So I'd say that for now the Altai-Sayan region looks like a pretty good option for the homeland of the ANE people (aka Mal'ta cluster).
Climatic changes during the Late Pleistocene had major impacts on the populations of many plant and animal species. During this period, humans and large mammals, e.g. brown bear and moose, were subject to analogous phylogeographic pressures and were linked by important ecological processes. However, evidence of human dispersal in northern Eurasia during the Late Pleistocene is scarce. Analysis of large mammals therefore has the potential to shed light on the population processes of humans during this period. We address several unresolved issues regarding the Late Pleistocene demography of brown bears: (a) the putative locations of refugia; (b) the direction of migrations across Eurasia and into North America; and (c) parallels with other mammals, including humans. We present results based on more than 200 complete mitochondrial genome sequences from Eurasian and North-American brown bears. Bayesian phylogenetic analysis revealed that most individuals belong to a very large Holarctic clade. The MRCA of this clade lived ca 40 thousand years ago, most likely in the Altai-Sayan area, a known Late Pleistocene refugium in Asia. We propose several migration scenarios for bears and suggest that brown bears and humans underwent a series of parallel migrations in Eurasia and to North America during the Late Pleistocene. Moreover, both species exhibited a demographic standstill in Beringia before colonizing North America. Synchrony in the timing of past migrations and standstill implies that the ecology of large mammals includes key limiting factors that can enhance our understanding of ancient human movements and on large carnivore conservation.Saarma et al., Parallel migrations in brown bears and humans within Eurasia and into North America during the Late Pleistocene, ConGenomics 2016 abstract.
Wednesday, May 18, 2016
Yamnaya = Khvalynsk + extra CHG + maybe something else
Thanks to recently published ancient DNA it's now generally accepted that the Yamnaya pastoralists of the Pontic-Caspian Steppe were basically a mixture of Eastern European Hunter-Gatherers (EHG) and Caucasus Hunter-Gatherers (CHG).
However, I thought it might be useful to dig a little deeper with the D-stats/nMonte method to see what else crops up. I put together a special datasheet for the job, with Yamnaya Samara featured both in the rows and columns. It's available for download here. The nMonte R script can be gotten here.
Using the most plausible reference samples currently available - almost all of them older than Yamnaya, and thus unlikely to skew the results with Yamnaya admixture - reveals the following models for the two Yamnaya sets from Kalmykia and Samara, respectively.
Yamnaya_Kalmykia Khvalynsk 57.7 Kotias 28.3 Hungary_EN 12.9 Ulchi 1.1 AfontovaGora3 0 Anatolia_Neolithic 0 Karelia_HG 0 Loschbour 0 MA1 0 Motala_HG 0 distance%=1.9125 / distance=0.019125 Yamnaya_Samara Khvalynsk 56.75 Kotias 26.4 Hungary_EN 10.85 Karelia_HG 4.4 Loschbour 1.6 AfontovaGora3 0 Anatolia_Neolithic 0 MA1 0 Motala_HG 0 Ulchi 0 distance%=2.1354 / distance=0.021354Very interesting but hardly surprising. Essentially what we're seeing there is potentially very strong genetic continuity from the Eneolithic to the Early Bronze Age on the Pontic-Caspian Steppe. In other words, from Khvalynsk to Yamnaya. However, at some point between the Eneolithic and the Early Bronze Age, the steppes saw a major influx of extra CHG, represented by the ~27% of Kotias-related admixture. Considering the relevant uniparental data, with lots of Y-HG R1b and no Y-HG J among Yamnaya males, I'd say this CHG came with women. Also, the relatively high admixture related to early Hungarian Plain farmers (Hungary EN) is a fairly curious detail that has not been reported before. If real, it probably represents gene flow from the Neolithic and/or Chalcolithic Balkans to the Pontic-Caspian Steppe. Again, in all likelihood it mostly came with women, perhaps from Tripolye-Cucuteni and/or Varna communities. But, to make sure the results weren't erroneous - perhaps skewed by the worryingly high affinity between the two CHG genomes, Kotias and Satsurblia, both pseudo-diploid sequences - I reran the tests using a different version of the same datasheet (available here). In this version, Kotias and Satsurblia are part of a single Caucasus HG sample included in the rows.
Yamnaya_Kalmykia Khvalynsk 53.95 Caucasus_HG 38.45 Hungary_EN 4.7 Motala_HG 2.9 AfontovaGora3 0 Anatolia_Neolithic 0 Karelia_HG 0 Loschbour 0 MA1 0 Ulchi 0 distance%=1.7492 / distance=0.017492 Yamnaya_Samara Khvalynsk 56.7 Caucasus_HG 33.55 Hungary_EN 4.25 Loschbour 2.95 Motala_HG 2.55 AfontovaGora3 0 Anatolia_Neolithic 0 Karelia_HG 0 MA1 0 Ulchi 0 distance%=2.0964 / distance=0.020964Yup, the Hungary EN contributions take a dive. But they're still above noise level (>2%), so they might well represent a real signal that entered the Yamnaya horizon or late Khvalynsk from the west. Can't wait to see how Yamnaya genomes from north of the Black Sea come out. Update 20/08/2017: It appears that I was onto something. See here. See also... Modeling Steppe_EMBA Yamnaya =/= Eastern Hunter-Gatherers + Iran Chalcolithic Another look at the genetic structure of Yamnaya
Saturday, May 14, 2016
D-stats/nMonte open thread #2
I put together a couple of D-stats datasheets featuring the Ice Age European samples from Qiaomei Fu et al. 2016. They're available for download here and here, and can be used as input for nMonte and 4Mix. For instance, a couple of models using nMonte:
Villabruna ElMiron 99.8 Anatolia_Neolithic 0.2 AfontovaGora3 0 Esan_Nigeria 0 GoyetQ116-1 0 Kostenki14 0 Kotias 0 MA1 0 Neandertal_Altai 0 Ulchi 0 Vestonice16 0 distance%=5.9112 / distance=0.059112 Kotias Anatolia_Neolithic 56.45 Kostenki14 24.1 AfontovaGora3 18.1 Esan_Nigeria 1.35 ElMiron 0 GoyetQ116-1 0 MA1 0 Neandertal_Altai 0 Ulchi 0 Vestonice16 0 Villabruna 0 distance%=8.3858 / distance=0.083858Update 17/05/2016: I wanted to see what would happen if I changed the formula from D(Chimp,Rows)(Mbuti,Columns) to D(Chimp,Columns)(Mbuti,Rows). The new datasheets are available here and here. The results are very similar, although never identical, and I'm not sure at the moment which set of sheets works better.
Villabruna ElMiron 99.5 Anatolia_Neolithic 0.5 AfontovaGora3 0 Esan_Nigeria 0 GoyetQ116-1 0 Kostenki14 0 Kotias 0 MA1 0 Neandertal_Altai 0 Ulchi 0 Vestonice16 0 distance%=5.8334 / distance=0.058334 Kotias Anatolia_Neolithic 69.5 AfontovaGora3 24.3 Ulchi 5.35 Kostenki14 0.85 ElMiron 0 Esan_Nigeria 0 GoyetQ116-1 0 MA1 0 Neandertal_Altai 0 Vestonice16 0 Villabruna 0 distance%=8.6196 / distance=0.086196See also... Yamnaya = Khvalynsk + extra CHG + maybe something else D-stats/nMonte open thread D-stats/4mix tour of ancient Eurasia
Thursday, May 12, 2016
Following the mammoth herds?
The recent Qiaomei Fu et al. paper on the population genomics of Ice Age Europe was a fascinating read, but its sampling strategy left a big blind spot: Eastern Europe 24-34 kyr BP. Check out what happens to the maternal lineages of Eastern European mammoths at this time.
Basically, they're replaced by lineages native to North America. During the 14-24 kyr BP period, the rest of the European mammoth population is also affected.
Of course, this is also precisely the period when the so called Villabruna hunter-gatherer cluster appears in Central Europe, probably as a result of mixture between the remnants of post-Ice Age Europeans and a population relatively closely related to Caucasus and Siberian hunter-gatherers (see the post and discussion here). Were these perhaps mammoth hunters?
No wonder then, that this is also the time when we see the first appearance of Y-chromosome haplogroup R1 in Europe; more precisely R1b1, in the ~14 kyr cal BP genome from Villabruna, northeast Italy, that defines the Villabruna cluster. After all, R is the sister clade of Q, the dominant Y-chromosome haplogroup in many parts of North Asia and the Americas.
Image credit...
Palkopoulou et al., Holarctic genetic structure and range dynamics in the woolly mammoth, Proc. R. Soc. B 2013 280 20131910; DOI: 10.1098/rspb.2013.1910. Published 11 September 2013
Friday, May 6, 2016
Villabruna cluster =/= Near Eastern migrants
I've been running a lot of Treemix analyses with the samples from the recent Qiaomei Fu et al. paper. And the impression I'm getting is that the authors missed the elephant in the room; the one with R1b painted on its big butt.
Now, it's true that Treemix output can't be used as unambiguous evidence in support of complex models. That's because in the absence of key samples the algorithm can get exceedingly creative in modeling the available data, sometimes to such extremes that the results might seem absurd.
However, when something keeps showing up again and again, even when using somewhat different samples and marker sets, and makes sense in the context of haploid and archaeological data, then at the very least it deserves serious consideration.
Here's a nice Treemix series that more or less captures the meat and potatoes of my many Treemix experiments with the Qiaomei Fu et al. dataset. For the archaeological contexts and other details about these ancient samples see here.
Below is my interpretation of the results:
- Villabruna is a sister clade of the earlier European Vestonice clade, but with significant input from an AfontovaGora3-related North Eurasian population, perhaps one that was living north of the Black Sea after the Kostenki people went the way of the dodo - Hence, the R1b lineage carried by Villabruna I9030, the individual in this Treemix series, probably comes from the Eurasian steppe - Kotias, a Caucasus Hunter-Gatherer, is in large part derived from the same North Eurasian population, hence the close relationship between Villabruna and Caucasus Hunter-Gatherers - Villabruna and/or closely related foragers contributed significant ancestry to Neolithic Anatolians, and thus, indirectly, possibly to all extant Near Eastern and even many African populations - Kotias is also closely related to Neolithic Anatolians, but probably mostly via a more basal population, perhaps the so called Basal Eurasians, native to the Near East prior to the Villabruna and/or related gene flow across the Near East and parts of Africa - Present-day East Asians might be ancient hybrids with admixture from the same or very similar North Eurasian population, although as per the above mentioned quirks of Treemix, it's possible that the North Eurasians that contributed ancestry to Villabruna, Caucasus foragers and Eurasian steppe populations were in fact partly East AsianSo basically what I'm seeing are back migrations from Europe and the Eurasian steppe or Siberia to the Near East soon after the Ice Age. Yes, from Europe, although I admit that things can get fuzzy here. That's because at the time much of the Aegean Sea was dry land, and thus there was no geographic barrier between the European Balkans and Asian Anatolia. The two regions, which might seem very distinct to us today, were basically one. So when I say Europe, I actually mean the ancient landmass now divided between the Balkans and Anatolia. Or not? Are there any D-stats that we can run to either confirm or debunk my Treemix-based hypothesis? Feel free to post your proposals in the comments and I'll try and run them as soon as possible. Update 07/05/2016: The plot thickens somewhat. I added MA1 to the line up, and now Villabruna shows minor Kotias-related ancestry. In other words, probably something from the Caucasus. The full series is available in a zip file here. However, although interesting, this result doesn't change much. In fact, it adds more weight to the argument that Villabruna inherited his Y-chromosome from steppe foragers. That's because if this migration edge was a reflection of gene flow from Anatolia to Europe, I'd expect it to run from the Anatolia Neolithic branch, rather than the Kotias branch. See also... Following the mammoth herds?
Thursday, May 5, 2016
On the modern genetic affinities of Ice Age Europeans
Apparently most of them didn't contribute much to our genomes. But for what it's worth, I tested their present-day affinities, and those of several other ancients, with f3 outgroup stats of the form f3(Test,X;Mbuti). For the archaeological contexts and other details about these ancient samples see here.
Not long ago Basques were thought to be by far the best proxies for Upper Paleolithic Europeans; almost a genetic relict from Ice Age Iberia. Clearly, this is not what they are, but they do share relatively high drift with the El Miron lady, which perhaps suggests some genetic continuity in Basque Country for almost 20,000 years. This would still be a remarkable story if confirmed via multiple lines of evidence.
Below are a couple of examples of what can be done with these stats. It's pretty clear that the Vestonice cave men show a preference for Northeast Europeans, in particular Balts, Estonians and Poles. On the other hand, Kostenki14 shares the highest drift with the Dutch, mostly north of the Rhine Dutch. Might that be meaningful in some way?
Are there any other interesting patterns in this data? Please note, however, that it's not possible to compare the stats directly, because they're based on different numbers of markers.
Wednesday, May 4, 2016
Ice Age Europeans in a global PCA
The datasheet can be downloaded here. It might be possible to model many of the populations in the PCA as mixtures of Ice Age Europeans and other ancients using nMonte, but I haven't tried this yet. Most of the samples in this run are from various Reich lab public data releases available here.
See also...
On the modern genetic affinities of Ice Age Europeans
PC/nMonte open thread
For a typical West Eurasian PCA, the Epipaleolithic is the final frontier
Tuesday, May 3, 2016
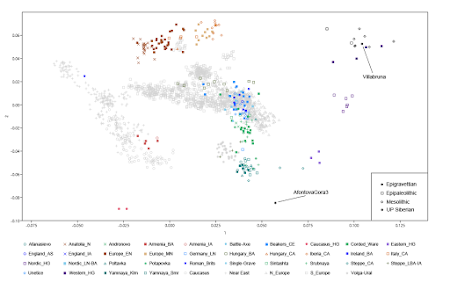
For a typical West Eurasian PCA, the Epipaleolithic is the final frontier
My dataset now includes the samples from Qiaomei Fu et al. 2016. They're freely available at the Reich lab website here.
Below is a Principal Component Analysis (PCA) featuring 10 of the freshly sequenced genomes from the paper classified as Epigravettian, Epipaleolithic and Mesolithic, as well as the Siberian (late?) Upper Paleolithic forager Afontova Gora 3.
I did try to include all of the new genomes from the paper on the plot, but most were getting results that didn't make any sense. They were simply too old and too far removed in terms of genetic structure from present-day West Eurasians. But that's OK. I can analyze them with formal statistics, and promise to do so vigorously in the coming weeks.
By the way, Villabruna is the ~14,000 year old genome from Italy belonging to Y-chromosome haplogroup R1b. Gioiello, you still breathing?
See also...
Ice Age Europeans in a global PCA
Monday, May 2, 2016
The genetic history of Ice Age Europe (Qiaomei Fu et al. 2016)
Just in at Nature:
Abstract: Modern humans arrived in Europe ~45,000 years ago, but little is known about their genetic composition before the start of farming ~8,500 years ago. Here we analyse genome-wide data from 51 Eurasians from ~45,000–7,000 years ago. Over this time, the proportion of Neanderthal DNA decreased from 3–6% to around 2%, consistent with natural selection against Neanderthal variants in modern humans. Whereas there is no evidence of the earliest modern humans in Europe contributing to the genetic composition of present-day Europeans, all individuals between ~37,000 and ~14,000 years ago descended from a single founder population which forms part of the ancestry of present-day Europeans. An ~35,000-year-old individual from northwest Europe represents an early branch of this founder population which was then displaced across a broad region, before reappearing in southwest Europe at the height of the last Ice Age ~19,000 years ago. During the major warming period after ~14,000 years ago, a genetic component related to present-day Near Easterners became widespread in Europe. These results document how population turnover and migration have been recurring themes of European prehistory.Fu et al., The genetic history of Ice Age Europe, Nature 02/05/2016; DOI:10.1038/nature17993 Here are three summaries of the findings from the comments:
Krefter: Europeans dating 36,000-30,000 years old from Russia, Belgium, Czech, and Italy form a clade with 24,000 year old Siberian MA1. But they're more related to each other than to MA1. Also, none of them have an especially close relationship to each other, unless they come from the same archaeological site/region. DNA from a lady who lived in Spain 19,000 years ago, has a close relationship to people who lived in Belgium 30,000 years ago. She also has a close relationship to "WHGs". So, she looks like a mixture of WHG and those 30,000 year old people from Belgium. After 14,000 years ago, Europeans from Northern Italy, Germany, France, Luxembourg, Spain, and Hungary all form a tight cluster. They are WHG. They also appear to have minor ancestry related to the 30,000 year old people who lived in Belgium. Stone age Middle Easterners CHG(Caucasus) and especially EEF(Turkey) have WHG-related ancestry. They're more related to post-14,000 years ago Europeans than to earlier Europeans. There's an unknown link between the Middle East and WHG. ... Matt: Trying to interpret this (and probably getting something wrong) the genetic history and cultural of Europe from this so far looks like this - 42,000-37,580 BP - Oase1 type people (and possibly others) living in Europe (Romania) - 35,160-34,430 BP - GoyetQ116 type people living in Europe (Belgium) (GoyetQ116 is classified to the Aurignacian culture wave). - 34,580-31,210 BP - Vestonice cluster types seem to replace the GoyetQ116 type (Except the GoyetQ116 type, Aurignancians(?) are likely to still be living in Iberia, as they're ancestral to the El Miron cluster, from later in history) Vestonice types seem to all be classified as Gravettian. - Glaciation then wipes out the Vestonice / Gravettian types around 24,000 BP. - 18,830-18,610 BP - El Miron is an admixture of a GoyetQ116 and the Villabruna branch (the latter thought to be Near Eastern in origin due to greater sharing with present day Near Eastern people) All samples from then on in Europe, are identified with the Magdalenian culture and with the same cluster membership as El Miron, as these guys expand across Europe after the retreat of the ice. - 14,180-13,780 BP - Villabruna branch ancestry in samples increases a lot, and material culture transitions to other industries - Epigravettian, Azilian, Epipaleolithic and Mesolithic. Looks like another great migration to Europe (out of the East via Italy?). ... Colin: There was no Basal Eurasian admixture in pre Neolithic Europe. The reasoning was that Ust Ishim (45kya Eurasian) does not contain Basal Eurasian and Ust Ishim is equally related to East Asians and pre Neolithic Europeans. There was no ANE, Mal'ta cluster, admixture in pre Neolithic Europe (presumably they are talking about the portion of Europe that is West of Russia and Ukraine). The reasoning here is that the mal'ta cluster is equally related to all pre Neolithic "Europeans" starting with Kostenki (37kya). -Consequently, ANE only entered "Europe" during the Neolithic and Bronze Age, probably from the Eurasian Steppe. So far, GoyetQ (35kya Belgium) and Kosteni (37kya Russia) are the first ancient Europeans to exhibit closer relations to modern Europeans over East Asians. The older Eurasian samples of Ust Ishim and Oaste do not show higher affinity to modern Europeans over East Asians. European samples from about 37kya to 14kya tend to be equally unrelated to non-European Eurasians. However, there are 14kyo samples, representing the Villabruna cluster, that show a higher affinity to the modern Near Easterners. The 13kyo to 10kyo samples from the Caucasus, representing the Satsurblia cluster, show more affinity to the Villabruna cluster than to the earlier Europeans. The Villabruna cluster can not be the result of admixture with the Satsurblia cluster since the former does not contain Basal Eurasian while the later does. GoyetQ (35kya Belgium) type ancestry shows a resurgence in the Miron Cluster of 19kya centered in Iberia. About half of the ancestry in the Miron Cluster derives from GoyetQ types. Because of the large number of individuals in the Villabruna cluster, the researchers were able to identify structure within the cluster. One interesting tid bit is that certain individuals in the Villabruna Cluster showed extra affinity towards East Asians. The oldest of which is the 13kyo sample from Switzerland. Two other Villabruna individuals, Loschbour and La Brana, for example, show extra affinity to Han Chinese over Kostenki. Because all three individuals are equally related to Ust Ishim, hypothetical differences in levels of Basal Eurasian cannot explain the greater affinity Loschbour and La Brana have to the Han.See also... On the modern genetic affinities of Ice Age Europeans For a typical West Eurasian PCA, the Epipaleolithic is the final frontier
Sunday, May 1, 2016
ESHG 2016 abstracts
See the Programme Planner here. Below is an abstract on the population history of the Caucasus. The emphasis is mine. Who actually believes the 20,000 year estimate? It sounds far fetched to me; probably blown out by minor East and/or South Asian admixture not accounted for by the authors.
Abstract: Archaeological findings suggest that modern humans have inhabited the Caucasus since Paleolithic times but little is known about their origin and how they have interacted with neighboring populations. Previous genetic studies of present-day populations from the Steppe suggested the Caucasus acted as a barrier, leading to genetic discontinuity between the Caucasus and the East European Plain. Furthermore, geography, ethnicity, and language are also expected to influence the mating patterns of the populations inhabiting the Caucasus and thus create genetic structure within this region. Here we use whole-genome genotyping and sequencing data to fully understand the genetic diversity of the Caucasus and its role in the demographic process that shaped modern Eurasian genomes. Combining data from 12 high-coverage whole-genome sequences, four each from Georgia, Armenia, and Azerbaijan with SNP-genotype data from more than 250 individuals from the same populations, we present one of the most comprehensive datasets for this region. Principal component analyses, followed by model-based clustering, reveal previously unreported structure among the Caucasus populations. In particular, we discovered three different clusters among Georgians and two in Azerbaijan, in addition to different ancestral proportions in these clusters. MSMC analysis of the whole genomes shows Caucasus populations have diverged in the last 20,000 years from other Eurasians and have experienced different demographic changes in recent times. We thus provide a detailed analysis of the past and present demography of the Caucasus which will be useful for understanding the genetic diversity of this region and the relationship with European populations.Mezzavilla et al., Genetic structure and demographic history of the Caucasus: insights from whole-genome sequences, ESHG EMPAG 2016 Presentation Abstract, P18.020D See also... Ancient Polish admixture in Denmark
Subscribe to:
Posts (Atom)