search this blog
Showing posts with label Celtic. Show all posts
Showing posts with label Celtic. Show all posts
Monday, February 21, 2022
The Pict
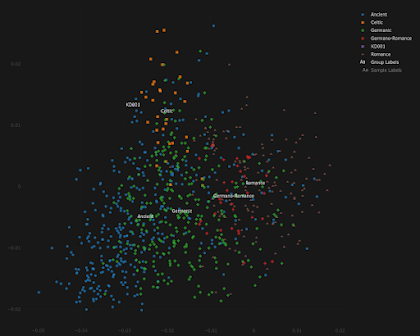
KD001 is the first undeniable Pictish sample in my dataset, courtesy of Dulias et al. 2022. Thanks to Altvred for processing the files.
This is how KD001 behaves in my Celtic vs Germanic Principal Component Analysis (PCA). Looks kind of Irish, doesn't he?
To see an interactive version of the plot, paste the coordinates from here into the relevant field here.
See also...
Celtic vs Germanic Europe
Avalon vs Valhalla revisited
When did Celtic languages arrive in Britain?
Labels:
Albion,
ancient DNA,
Britain,
British DNA,
British Isles,
Celtic,
Celtic vs Germanic,
Great Britain,
Indo-European,
Northern Europe,
Northwestern Europe,
Pict,
Pictish,
Picts,
Scotland,
Scottish
Thursday, December 23, 2021
When did Celtic languages arrive in Britain?
A new paper at Nature by Patterson et al. argues that Celtic languages spread into Britain during the Bronze Age rather than the Iron Age [LINK]. This argument is based on the observation that there was a large-scale shift in deep ancestry proportions in Britain during the Bronze Age.
In particular, the ratio of Early European Farmer (EEF) ancestry increased significantly in what is now England during the Late Bronze Age (LBA). On the other hand, the English Iron Age was a much more stable period in this context.
I don't have any strong opinions about the spread of Celtic languages into Britain, and Patterson et al. might well be correct, but their argument is potentially flawed because:
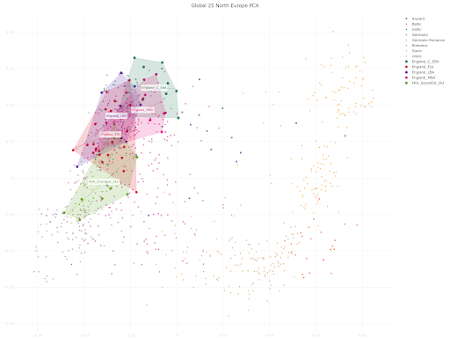
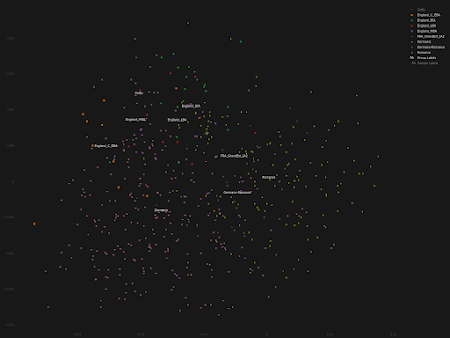
- significant population shifts need not result in any noticeable changes in ancient ancestry proportions - ancient ancestry proportions can shift without significant migrations from afar due to cryptic population substructures - large-scale population shifts need not result in langage shifts, especially if they're gradual - small-scale population shifts can result in language shifts, especially if they're sudden.Indeed, when I plot some of the key ancient samples from the paper in my ultra fine scale Principal Component Analyses (PCA) of Northern and Western Europe, it appears that it's only the Early Iron Age (EIA) population from England that overlaps significantly with a roughly contemporaneous group from nearby Celtic-speaking continental Europe. The relevant PCA data are available here and here, respectively. See also... Celtic vs Germanic Europe Avalon vs Valhalla revisited R1a vs R1b in third millennium BCE central Europe
Friday, August 27, 2021
R1a vs R1b in third millennium BCE Central Europe (Papac et al. 2021)
R1a-M417 and R1b-L51 are by far the most important Y-chromosome haplogroups in Europe today. More precisely, R1a-M417 dominates in Eastern Europe, while R1b-L51 in Western Europe.
It's been obvious for a while now, at least to me, that both of these Y-haplogroups are closely associated with the men of the Late Neolithic Corded Ware culture (CWC). Indeed, in my mind they're the main genetic signals of its massive expansion, probably from a homeland somewhere north of the Black Sea in what is now Ukraine.
I'm still not exactly sure how the east/west dichotomy between R1a and R1b emerged in Europe, but, thanks to a new paper by Papac et al. at Science Advances, at least now I have a working hypothesis about that. Below is a quote from the said paper, emphasis is mine:
In addition to autosomal genetic changes through time, we observe a sharp reduction in Y-chromosomal diversity going from five different lineages in early CW to a dominant (single) lineage in late CW (Fig. 4A). We used forward simulations to explore the demographic scenarios that could account for the observed reduction in Y-chromosomal diversity. Performing 1 million simulations of a population with a starting frequency of R1a-M417(xZ645) centered around the observed starting frequency in Bohemia_CW_Early (3 of 11, 0.27), we assessed the plausibility of this lineage reaching the observed frequency in Bohemia_CW_Late (10 of 11, 0.91) in the time frame of 500 years under a model of a closed population and random mating (Materials and Methods). We reject the “neutral” hypothesis, i.e., that this change in frequency occurred by chance, given a wide range of plausible population sizes. Instead, our results suggest that R1a-M417(xZ645) was subject to a nonrandom increase in frequency, resulting in these males having 15.79% (4.12 to 44.42%) more surviving offspring per generation relative to males of other Y-haplogroups. We also find that this change in Y chromosome frequency is extreme compared to the changes in allele frequencies at fully covered autosomal 1240k sites within the same males, suggesting a process that disproportionately affected Y-chromosomal compared to autosomal genetic diversity, ruling out a population bottleneck as the likely cause. Our results suggest that the Y-lineage diversity in early CW males was supplanted by a nonrandom process [selection, social structure, or influx of nonlocal R1a-M417(xZ645) lineages] that drove the collapse in Y-chromosomal diversity. A simultaneous decline of Y-chromosomal diversity dating to the Neolithic has been observed across most extant Y-haplogroups (64), possibly due to increased conflict between male-mediated patrilines (65). We view that changes in social structure (e.g., an isolated mating network with strictly exclusive social norms) could be an alternative cause but would be difficult to distinguish in the underlying model parameters.Right, so even though the CWC was clearly a community of closely related groups, there must have been some competition between its different clans. And since these clans were highly patriarchal and patrilineal, this competition probably led to different paternal lineages dominating different parts of the CWC horizon, with M417 becoming especially common in the east and L51 in the west. Of course, the expansions of post-Corded Ware groups, such as the M417-rich Slavs in Eastern Europe and L51-rich Celts in Western Europe, were also instrumental in creating Europe's R1a/R1b dichotomy, but obviously these groups were in large part the heirs of the CWC. By the way, most of the samples from Papac et al. are already in the Global25 datasheets linked here. Look for the labels listed here. Below is a plot made from the Global25 data courtesy of regular commentator Matt. Citation: L. Papac, M. Ernée, M. Dobeš, M. Langová, A. B. Rohrlach, F. Aron, G. U. Neumann, M. A. Spyrou, N. Rohland, P. Velemínský, M. Kuna, H. Brzobohatá, B. Culleton, D. Daněček, A. Danielisová, M. Dobisíková, J. Hložek, D. J. Kennett, J. Klementová, M. Kostka, P. Krištuf, M. Kuchařík, J. K. Hlavová, P. Limburský, D. Malyková, L. Mattiello, M. Pecinovská, K. Petriščáková, E. Průchová, P. Stránská, L. Smejtek, J. Špaček, R. Šumberová, O. Švejcar, M. Trefný, M. Vávra, J. Kolář, V. Heyd, J. Krause, R. Pinhasi, D. Reich, S. Schiffels, W. Haak, Dynamic changes in genomic and social structures in third millennium BCE central Europe. Sci. Adv. 7, eabi6941 (2021). See also... On the origin of the Corded Ware people Understanding the Eneolithic steppe Conan the Barbarian probably belonged to Y-haplogroup R1a
Labels:
ancient DNA,
Balto-Slavic,
Celtic,
Central Europe,
Corded Ware,
Corded Ware Culture,
CWC,
Eastern Europe,
Germanic,
Indo-European,
Northern Europe,
Proto-Indo-European,
R1a-M417,
R1b-L151,
R1b-L51,
Slavic,
Slavs
Sunday, January 17, 2021
That old chestnut: Northeast vs Northwest Euros
In the last comment thread reader Greg put forth this question:
David, when are you going to explain the genetic discrepancy between Northeastern and Northwestern Europeans? You know, the one that people believe is due to Baltic Hunter-Gatherer admixture, whereas you believe it is due to genetic drift? You ought to make a post about this issue at some point, because a lot of people are wondering what's causing the differences.Well, Greg, this issue has been discussed to the proverbial death here and elsewhere. In fact, there were two posts and rather lengthy comment threads on the same topic at this blog just a few months ago. See here and here. Nevertheless, it seems that a fair number of people are still befuddled, so I'm going to try to explain this one last time, as briefly as a I can using just a handful of f4-stats. Admittedly, Northeast Europeans generally do pack higher levels of indigenous European hunter-gatherer ancestry than Northwest Europeans. This is especially true of Balts, who show more of this type of ancestry than even Scandinavians in practically every type of analysis. The f4-stats below back this up unambiguously. Note the significantly positive (>3) Z scores, which suggest that Latvians and Lithuanians harbor more Baltic hunter-gatherer-related ancestry than Norwegians and Swedes.
Chimp Baltic_HG Norwegian Latvian 0.001301 7.114 Chimp Baltic_HG Swedish Latvian 0.001017 4.205 Chimp Baltic_HG Norwegian Lithuanian 0.001023 7.341 Chimp Baltic_HG Swedish Lithuanian 0.000763 3.408Greg, I know what you're thinking: the naysayers are right! But wait, because there's a twist to this tale. Check out these f4-stats:
Chimp Baltic_HG Norwegian Belarusian 0.000265 1.934 Chimp Baltic_HG Swedish Belarusian 0.000152 0.7 Chimp Baltic_HG Norwegian Polish 6.4E-05 0.519 Chimp Baltic_HG Swedish Polish -0.000235 -1.074Please note, Greg, that none of the Z scores reach significance, which means that these Northwest Europeans and Slavs are symmetrically related to Baltic_HG. They're also symmetrically related to other relevant ancient groups such as the Yamnaya steppe herders. This, of course, suggests that they harbor very similar levels of basically the same ancient genetic components.
Chimp Karelia_HG Norwegian Belarusian 0.000136 0.844 Chimp Karelia_HG Swedish Belarusian 7.9E-05 0.32 Chimp Karelia_HG Norwegian Polish -4.7E-05 -0.304 Chimp Karelia_HG Swedish Polish -0.000134 -0.54 Chimp Yamnaya_Samara Norwegian Belarusian -0.000134 -1.085 Chimp Yamnaya_Samara Swedish Belarusian -6.6E-05 -0.34 Chimp Yamnaya_Samara Norwegian Polish -0.000225 -1.995 Chimp Yamnaya_Samara Swedish Polish -0.000311 -1.574 Chimp Barcin_N Norwegian Belarusian -0.000335 -2.809 Chimp Barcin_N Swedish Belarusian -0.000284 -1.491 Chimp Barcin_N Norwegian Polish -0.000222 -2.057 Chimp Barcin_N Swedish Polish -0.000318 -1.662 Chimp Baikal_N Norwegian Belarusian 0.000186 1.3 Chimp Baikal_N Swedish Belarusian -7E-05 -0.33 Chimp Baikal_N Norwegian Polish -4.6E-05 -0.351 Chimp Baikal_N Swedish Polish -0.000477 -2.277Interestingly, pairing up Ukrainians with English samples from Cornwall and Kent produces similar outcomes. But that's because most ancient ancestry proportions in Europe show a closer correlation with latitude than longitude.
Chimp Baltic_HG English_Cornwall Ukrainian 0.000282 2.242 Chimp Baltic_HG English_Kent Ukrainian 0.000225 1.748 Chimp Karelia_HG English_Cornwall Ukrainian 0.000323 2.175 Chimp Karelia_HG English_Kent Ukrainian 0.000239 1.634 Chimp Yamnaya_Samara English_Cornwall Ukrainian -6.6E-05 -0.569 Chimp Yamnaya_Samara English_Kent Ukrainian -0.000112 -0.977 Chimp Barcin_N English_Cornwall Ukrainian -0.000519 -4.641 Chimp Barcin_N English_Kent Ukrainian -0.000598 -5.232 Chimp Baikal_N English_Cornwall Ukrainian 0.000385 2.874 Chimp Baikal_N English_Kent Ukrainian 0.00036 2.836Now, Greg, if at least in terms of genetic ancestry, Latvians, Lithuanians, Belarusians, Poles and Ukrainians all qualify as Northeast Europeans, then what makes them different, as a group, from Northwest Europeans? Do you believe that the key factor is admixture from Baltic hunter-gatherers? Or is it genetic drift? Of course, considering all of the f4-stats above, logic dictates that it must be relatively recent genetic drift. Keep in mind, however, that this only applies to Balto-Slavic speaking Northeast Europeans without significant Uralian ancestry. Overall, Uralic speakers have a more complex population history, and indeed genetic differences between them and Northwest Europeans are in large part due to somewhat different ancestry proportions and also Siberian admixture. See also... So who's the most (indigenous) European of us all?
Tuesday, April 21, 2020
Aesch25
During the early 3rd Millennium BC much of Central and Northern Europe was being infiltrated by pioneer herders, often young men, from the east associated with the Corded Ware culture (CWC).
In some important ways, this expansion may have been very similar to the European colonization of the more remote parts of the Americas during the 16th and 17th centuries.
For instance, the European newcomers weren't always able to dominate the indigenous peoples, and, sometimes, instead of trying to impose their culture on them, they accepted theirs.
I suspect that Aesch25, an ancient sample from the recent Furtwängler et al. paper on the social and genetic structure of the prehistoric populations of the Swiss Plateau, represents a similar case.
Aesch25 wasn't buried with grave goods so he wasn't given a cultural context in the said paper. However, dated to 2864-2501 calBC, he's the earliest individual in this part of Europe with the originally Eastern European Y-haplogroup R1b-M269 and a CWC-like genome-wide genetic structure.
Indeed, the other fourteen samples from the same burial site, dated to more or less the same period as Aesch25, are overwhelmingly of local Neolithic farmer origin.
In any case, irrespective of his cultural affiliation and life story, Aesch25 represents an important data point in the search for the homeland of the so called Bell Beakers who spread across much of Europe during the Copper Age. That's because most Bell Beaker males belong to R1b-M269 and are very similar to Aesch25 in terms of overall genetic structure, apart from an excess of Neolithic farmer ancestry.
My view is that the Bell Beakers were an offshoot of the Single Grave culture (SGC), the westernmost variant of CWC. Of course, the SGC was centered on what is now northwestern Germany and surrounds, and didn't reach into the Swiss Plateau. However, in all likelihood it was founded by men closely related to Aesch25.
Below is a Principal Component Analysis (PCA) based on Global25 data featuring Aesch25 and several other individuals from the Furtwängler et al. paper. To view an interactive version of the plot, copy paste the data from the text file here into the relevant field here, then press Add to PCA. Also, you should copy paste each population separately to make sure that they don't form one grouping in the PCA key.
Aesch25 can easily pass for a CWC individual from what is now Germany (DEU_CWC_LN). On the other hand, the CWC samples from the Swiss Plateau (CHE_CWC_LN) are clearly shifted "south" relative to the German CWC cluster, which suggests that they harbor more Neolithic farmer ancestry. Indeed, they all belong to Y-haplogroup I2, which is especially closely associated with Middle Neolithic European farmers.
MX265, from Singen in southwest Germany, is the only sample in the Furtwängler et al. dataset that belongs to Y-haplogroup R1a. This is a somewhat unexpected outcome, because R1a is, overall, the most common Y-haplogroup in CWC males (see here).
Another surprise is that this individual is dated to just 763-431 calBC, which is a period that overlaps with the Hallstatt and La Tene cultures in Central Europe. Considering that these cultures are often associated with early Celts, was this person perhaps the speaker of a long lost Celtic language?
See also...
Single Grave > Bell Beakers
Dutch Beakers: like no other Beakers
Hungarian Yamnaya > Bell Beakers?
Hungarian Yamnaya predictions
The Battle Axe people came from the steppe
Saturday, December 14, 2019
Avalon vs Valhalla revisited
Pictured below is a new version of my Celtic vs Germanic genetic map. It's based on the same Principal Component Analysis (PCA) as the original (which can be seen here), but more focused on Northwestern Europe and produced with a different program.
To see the interactive online version, navigate to Vahaduo Custom PCA and copy paste the text from here into the empty space under the PCA DATA tab. Then press the PLOT PCA button under the PCA PLOT tab. For more guidance, refer to the screen caps here and here.
To include a wider range of populations in the key, just edit the data accordingly. For instance, to break up the ancient grouping into more specific populations, delete the Ancient: prefix in all of the relevant rows. This is what you should see:
Conversely, you can leave the ancient sample set intact and instead reorder the present-day linguistic groupings into, say, geographic groupings. To achieve this just delete all of the linguistic prefixes, such as Celtic:, Germanic:, and so on. You should end up with a datasheet like this and plot like this.
Of course, you can design your own plot by using any combination of the ancient and present-day individuals and populations that I've already run in this PCA. Their coordinates are listed here. Indeed, if you're in the possession of your own Celtic vs Germanic PCA coordinates, you can add yourself to the plot. And if you're not, see here.
It's also possible to re-process PCA data via the SOURCE tab. But I don't recommend doing this with the Celtic vs Germanic data, which are derived from a fine scale analysis and don't pack much variation. On the other hand, Global25 data are ideal for such re-processing. I made the plots below from subsets of Global25 coordinates available in a zip file here. To see how, refer to the screen caps here and here.
See also...
Modeling your ancestry has never been easier
Getting the most out of the Global25
Modeling genetic ancestry with Davidski: step by step
Labels:
ancient DNA,
Anglo-Saxon,
Avalon,
British Isles,
Celtic,
Celtic vs Germanic,
Gaelic,
Germanic,
Ireland,
Irish,
Nordic,
North Sea,
Northern Europe,
Northwestern Europe,
PCA,
Scandinavia,
Valhalla,
Viking
Monday, November 25, 2019
Viking Age Iceland
I finally managed to get some of the Icelandic ancients from Ebenesersdóttir et al. 2018 into the Global25 datasheets (see here). Better late than never. Look for the"ISL_Viking_Age" prefix. Below is a screen cap of a Principal Component Analysis (PCA) with the new samples. It was done with an online Global25 PCA runner freely available here.
The individuals classified as unadmixed Gaels and Norse by Ebenesersdóttir et al. generally also look like it based on their Global25 coordinates.
The mixture models below, using all of the populations from the Global25 "modern pop averages scaled" datasheet, were run with an online tool freely available here. Note that the ADD DIST COL option is set to 1X. This is a useful feature for modeling the fine scale ancestry of samples that are derived from very similar populations.
See also...
They came, they saw, and they mixed
Commoner or elite?
Who were the people of the Nordic Bronze Age?
Labels:
ancient DNA,
Bjork,
Celtic,
Gaelic,
Gaels,
Germanic,
Global25,
I1,
Iceland,
Icelandic,
Ireland,
Norse,
Norsemen,
Northern Europe,
PCA,
R1a-Z284,
R1b-L21,
Scandinavia,
Viking Age,
Vikings
Monday, March 25, 2019
Celtic probably not from the west
The term "Celtic from the west" is the catchphrase for a working theory, offered in a couple of recent books, positing that the earliest speakers of Celtic languages lived in Atlantic Europe during the Bronze Age or even earlier. It'll be interesting to see how this theory holds up against increasing numbers of ancient samples from attested early Celtic-speaking populations.
More popular and long-standing theories postulate that the Proto-Celts are associated with the Urnfield and/or Hallstatt archeological cultures of Late Bronze Age and Iron Age Central Europe. I'm inclined to agree with these more mainstream views when looking at my qpAdm mixture models below of three Celtiberians from what is now La Hoya, northern Spain, from the recent Olalde et al. paper on the genomic history of Iberia.
Celtiberian_LaHoya Halberstadt_LBA 0.207±0.077 Pre-Celtiberian_LaHoya 0.793±0.077 chisq 15.031 tail prob 0.522396 Full output Celtiberian_LaHoya Halberstadt_LBA 0.196±0.074 Non-Celtic_Iberian 0.804±0.074 chisq 17.366 tail prob 0.362297 Full outputThe Celtiberians show a stronger signal of (Urnfield-related?) ancestry from the northeast than their Bronze Age predecessors in northern Iberia (Pre-Celtic_LaHoya) as well as their Iron Age contemporaries from eastern Iberia (Non-Celtic_Iberian). The latter group very likely spoke the non-Indo-European Iberian language. It's not clear what the Bronze Age northern Iberians spoke, but it may have been a language related to Basque, which is also non-Indo-European. Of course, the fact that the Celtiberians harbored more northern Bell Beaker-related ancestry than basically all earlier Iberian groups was already reported in the Olalde et al. paper (on page 2), but I just wanted to see if I could flesh out some more details in regards to this observation by using chronologically and archeologically more proximate reference populations. See also... Open thread: What are the linguistic implications of Olalde et al. 2019? An exceptional burial indeed, but not that of an Indo-European Late PIE ground zero now obvious; location of PIE homeland still uncertain, but...
Labels:
ancient DNA,
Atlantic Bronze Age,
Atlantic Europe,
Basque,
Bell Beaker,
Celtiberian,
Celtic,
Hallstatt culture,
Iberia,
Iberian languages,
Indo-European,
Iron Age,
qpAdm,
Tartessian,
Urnfield culture,
Vasconic
Monday, March 18, 2019
Open thread: What are the linguistic implications of Olalde et al. 2019?
I was going to write a huge post on the linguistic implications of the latest batch of ancient DNA from Iberia courtesy of Olalde et al. 2019, and then I thought better of it. Admittedly, I don't know enough about the languages of prehistoric Iberia to say anything really useful on the topic. So instead here's an open thread to bounce around a few ideas in the comments.
Just briefly, this is what Olalde et al. say in the abstract of their paper about the relationship between ancestry from the Pontic-Caspian steppe and languages in Iron Age Iberia:
We reveal sporadic contacts between Iberia and North Africa by ~2500 BCE and, by ~2000 BCE, the replacement of 40% of Iberia’s ancestry and nearly 100% of its Y-chromosomes by people with Steppe ancestry. We show that, in the Iron Age, Steppe ancestry had spread not only into Indo-European–speaking regions but also into non Indo-European–speaking ones, and we reveal that present-day Basques are best described as a typical Iron Age population without the admixture events that later affected the rest of Iberia.However, in the paper it's revealed that "Indo-European regions" actually refers to a Celtic-speaking part of northern Iberia. And it's quite possible that Celts moved into this area from outside of Iberia only during the Iron Age. In other words, the speakers of Indo-European languages here may not have been the descendants of any of the people with steppe ancestry who came to Iberia by ~2000 BCE. So I'm probably not alone in thinking that the question of the linguistic affinities of these early migrants with steppe ancestry to Iberia (mostly associated with the Bell Beaker culture or BBC) remains open, especially since they evidently had such a profound genetic impact on the later non Indo-European-speaking populations of eastern and southern Iberia. Could they have been the speakers of unattested Indo-European languages, as well as Proto-Iberian and Proto-Basque? If not, why not? Below is a Principal Component Analysis (PCA) of West Eurasian genetic variation. I highlighted some of the ancient samples from Olalde et al., as well as Basques and other present-day Iberians. The Basques form a tight cluster with most of the Copper, Bronze and Iron Age Iberians, and, unlike the other present-day Iberians, they basically look like an Iberian population from the metal ages. The relevant datasheet is available here. This is nothing new and very much in line with the results in Olalde et al., but I wanted to emphasize the point that Basques were not just a group that experienced an extreme founder effect in R1b-P312, which is a Beaker-specific Y-chromosome lineage. Rather, they're still very similar to Iberian Beakers in terms of overall genetic structure. So where did they get their language? See also... Celtic probably not from the west Late PIE ground zero now obvious; location of PIE homeland still uncertain, but...
Labels:
ancient DNA,
Basques,
Celtic,
Celtic not from the west,
Hallstatt culture,
Iberia,
Iberian languages,
Indo-European,
Proto-Basque,
R1b-DF37,
R1b-L51,
R1b-P312,
Tartessian,
Urnfield culture,
Vasconic
Sunday, September 16, 2018
Celtic vs Germanic Europe
I have a feeling that ancient DNA from post-Bronze Age Northwestern Europe will be coming thick and fast from now on. To get the most out of such data I've designed a new Principal Component Analysis (PCA) that does a better job of separating the Celtic- and Germanic-speaking populations of Europe than my previous efforts of this sort (see here and here). Below are two different versions of the same PCA. The relevant datasheet is available here.
And here's a Discrimination Analysis (LDA) plot based on the 25 principal components. It further differentiates many of the populations along the east > west cline of genetic diversity.
The difference between the Germanic Anglo-Saxons and the Celtic and Roman Britons of what is now eastern England is obvious. The Anglo-Saxons could pass for Scandinavians, while the Celts and Romans both cluster between the Irish and French. This makes good sense, and is exactly what I was looking for. It's also interesting to see the presumably Celtic-speaking Hallstatt samples from Bylany, Czechia, clustering with the Belgians.
Update 14/12/2019: Pictured below is a new version of my Celtic vs Germanic genetic map. It's based on the same Principal Component Analysis (PCA) as the original, but more focused on Northwestern Europe and produced with a different program.
To see the interactive online version, navigate to Vahaduo Custom PCA and copy paste the text from here into the empty space under the PCA DATA tab. Then press the PLOT PCA button under the PCA PLOT tab. For more guidance, refer to the screen caps here and here.
To include a wider range of populations in the key, just edit the data accordingly. For instance, to break up the ancient grouping into more specific populations, delete the Ancient: prefix in all of the relevant rows. This is what you should see:
Conversely, you can leave the ancient sample set intact and instead reorder the present-day linguistic groupings into, say, geographic groupings. To achieve this just delete all of the linguistic prefixes, such as Celtic:, Germanic:, and so on. You should end up with a datasheet like this and plot like this.
Of course, you can design your own plot by using any combination of the ancient and present-day individuals and populations that I've already run in this PCA. Their coordinates are listed here. Indeed, if you're in the possession of your own Celtic vs Germanic PCA coordinates, you can add yourself to the plot. And if you're not, see here.
It's also possible to re-process PCA data via the SOURCE tab. But I don't recommend doing this with the Celtic vs Germanic data, which are derived from a fine scale analysis and don't pack much variation. On the other hand, Global25 data are ideal for such re-processing. I made the plots below from subsets of Global25 coordinates available in a zip file here. To see how, refer to the screen caps here and here.
See also...
Modeling your ancestry has never been easier
Getting the most out of the Global25
Modeling genetic ancestry with Davidski: step by step
Labels:
ancient DNA,
Anglo-Saxon,
Avalon,
British Isles,
Celtic,
Celtic vs Germanic,
Gaelic,
Germanic,
Ireland,
Irish,
Nordic,
North Sea,
Northern Europe,
Northwestern Europe,
PCA,
Scandinavia,
Valhalla,
Viking
Subscribe to:
Posts (Atom)