PL046 R-YP6228 PL048 I-PH833 PL049 I-A11537 PL052 R-Y48961 PL059 I-PH833 PL062 I-S15301 PL065 I-Y294193 PL066 R-FGC2555 PL067 R-S7759 PL070 I-CTS10028 PL071 I-BY316 PL076 I-S9318 PL082 I-Z2041 PL085 J-Z38241 PL086 I-FT29339See also... Early Slavs from Tribal Period Poland Wielbark Goths were overwhelmingly of Scandinavian origin High-resolution stuff
search this blog
Saturday, January 17, 2026
New Iron Age samples from southeastern Poland
Saturday, September 6, 2025
Early Slavs from Tribal Period Poland
The only deviation from this pattern is observed for ancient samples from the Russian Volga-Oka region, where we measure higher genetic affinity between present-day Southern/Western Europeans and the SP population compared to the pre-SP population (Fig. S17). This agrees with the pattern observed in PCA and ADMIXTURE that, in contrast to the Northwestern Balkan, Eastern Germany, and Poland-Northwestern Ukraine, the arrival of Slavic-associated culture in Northwestern Russia was associated with a shift in PCA space to the West, a decrease of BAL [Baltic] ancestry, and the introduction of Western European ancestries such as CNE [Continental North European] and CWE [Continental Western European].Thus, it's highly plausible that the Tribal Period Slavs from Gródek were very similar, perhaps even practically identical, to the proto-Slavs who lived in the original Slavic homeland. Hopefully we won't have to wait too long to discover whether that's true or not. More Migration period and Slavic period samples from the border regions of Belarus, Poland and Ukraine are needed to sort that out. On the other hand, most of the post-1000 CE individuals from Gródek are shifted closer to present-day Balts. This is probably due to admixture from nearby Baltic-speaking populations. At the time, Baltic speakers still occupied much of northern and eastern Poland. I'm still going through the Gretzinger et al. paper and I'll probably have a lot more to say about it in the near future. However, unfortunately, I've already spotted a silly mistake in the supplementary info that will probably have some very annoying consequences for us on this blog. On page 109 the authors make the false claim that South Asian ancestry is present in a wide range of ancient Eastern European and Central Asian populations from the Bronze Age to the Scythian period.
Furthermore, Sycthian groups from Ukraine show varying fractions of South Asian ancestry (between 5% and 12%), a component present in many ancient individuals from Moldova (e.g. Moldova_IA, Moldova_LBA and Moldova_MBA), Ukraine (Ukraine_Alexandria_MBA and Ukraine_BA_Catacomb.SG), Western Russia (e.g. Russia_EarlySarmatian.SG, Russia_MLBA_Potapovka, or Russia_MLBA_Sintashta) and the Caucasus (Russia_Caucasus_LBA_Dolmen and Russia_North_Caucasus_MBA) but (nearly) absent in the SP genomes from Central and East-Central Europe (<5%) (Fig. S42b).All ancient and present-day South Asian populations carry what is commonly known as Ancestral South Indian (ASI) ancestry, while all of the above mentioned ancient groups lack it. Ergo, it's impossible for these ancients to have actual South Asian ancestry. What happened is that Gretzinger et al. created a genetic component in ADMIXTURE based on present-day South Asians. However, South Asians today have very complex ancestry from several different sources, including early pastoralists from the North-Pontic steppe in Eastern Europe and early farmers from Central Asia and what is now Iran. As a result, the groups that share significant amounts of alleles with South Asians via these sources also show so called South Asian ancestry in the Gretzinger et al. analysis. Unless this problem is corrected we're likely to see some nutjobs online using this paper to claim all sorts of nonsense about the origins of ancient Eastern Europeans and Central Asians, especially the Sintashta people and Scythians. See also... High-resolution stuff Leo Speidel & Pontus Skoglund
Monday, March 25, 2024
High-resolution stuff
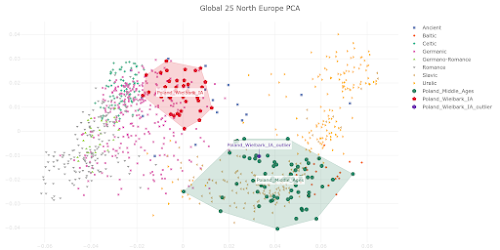
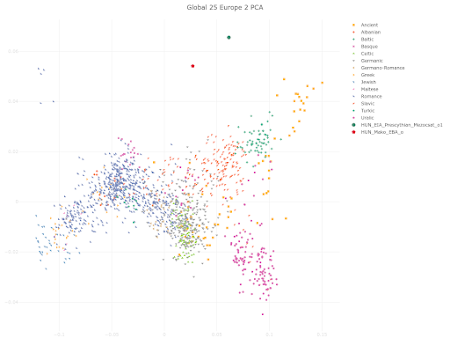
I appreciate that Polish population history is not the main focus of your preprint, and also that you're constrained by the lack of relevant and suitably high quality ancient genomes from East-Central and Eastern Europe. However, I must say that your analysis of the Medieval Polish population and resulting conclusions about Polish population history don't reflect reality. Your Poland_Middle_Ages genomic cluster is made up of just six samples that don't fully represent the genetic complexity of the core population of Medieval Poland. As a result, you classified PCA0148 as one of the Poland_Middle_Ages outliers, even though this sample isn't an outlier when analyzed within the context of the full set of published Polish Medieval genomes. Moreover, PCA0148 is very similar to several Polish Viking Age samples that show Scandinavian-specific genome-wide and Y-chromosome haplotypes, and probably likewise shows some Scandinavian-related ancestry. This is important to note when attempting to recapitulate Polish population history, because it suggests that Scandinavian-related ancestry played a formative role in the shaping of the core Polish Medieval genetic cluster. Thus, you might be correct when you claim that the six samples in your Poland_Middle_Ages cluster don't show any "detectable" Scandinavian-related ancestry, but this doesn't necessarily mean that this type of ancestry isn't a key part of the post-Iron Age Polish population history. Below is a self-explanatory Principal Component Analysis (PCA) plot that illustrates my points. Interestingly, Figure 3c in your preprint shows very similar outcomes in regards to the post-Iron Age Polish population history. But the style and scale of your figure makes it difficult to spot the subtle but likely genuine Northwest European-related genetic shifts shown by PCA0148, the Viking context samples and present-day Poles relative to the Poland_Middle_Ages cluster. However, I'm also skeptical that your Poland_Middle_Ages cluster doesn't carry any detectable or even significant Scandinavian-related ancestry. That's because I suspect that there might be some technical issues with your analysis that are masking this type of ancestry in the Polish samples. Your top mixture model for the Poland_Middle_Ages cluster is, in all likelihood, an extreme statistical abstraction of reality, rather than a close reflection of it. That's because, due to a combination of historical, geographical and genetic factors, neither Italy.Imperial(I).SG nor Lithuania.IronRoman.SG are realistic formative source populations for the Medieval Polish gene pool. One of the reasons why you ended up with such a surprising result is probably the lack of suitable samples from East-Central and Eastern Europe, especially those associated with plausibly the earliest Slavic-speaking populations. It's also possible that basing your mixture model on formal statistics played a key part. Formal statistics-based mixture models are known to be biased towards outcomes involving mixture sources from the extremes of mixture clines. If your analysis is affected by this problem, then this would help to explain why you characterized the Poland_Middle_Ages cluster as simply a two-way mixture between a Middle Eastern-related group from Imperial Rome and a Baltic population with a very high cut of European hunter-gatherer ancestry. I do note that on page 6 of your manuscript you consider the possibility that the Southern European-related signal in the Poland_Middle_Ages cluster might only be very distantly related to Italy.Imperial(I).SG, and that it may even have spread across Poland with early Slavic speakers. This is a great point, and I think it should be emphasized and expanded upon, because I suspect that the problem runs deeper than this. For instance, if the early Slavic ancestors of Poles carried substantially more Southern European-related ancestry than Lithuania.IronRoman.SG, and this ancestry was, say, more Balkan-related than Italian-related, then this might radically change your modeling of the Poland_Middle_Ages cluster. That's because these early Slavs would be positioned in a very different genetic space than Lithuania.IronRoman.SG, which could potentially require a significant signal of Scandinavian-related ancestry to get a robust mixture model. Finally, it might be useful to consider Isolation-by-Distance as a partial vector for the Italy.Imperial(I).SG-related signal in Medieval Poland. The full set of published Polish Medieval genomes includes a number of outliers with obvious ancestry from Western Europe and the Balkans. These people probably don't represent any large-scale migrations into Poland, but rather the movements of individuals and small groups. Over time, such small-scale mobility may have had a fairly significant impact on the genetic character of the Polish population.Update 26/03/2024: I sent another email to Speidel et al., this time in regards to their analysis of present-day Hungarians.
Your preprint also claims that present-day Hungarians are genetically similar to Scythians, and that this is consistent with the arrival of Magyars, Avars and other eastern groups in this part of Europe. However, present-day Hungarians are overwhelmingly derived from Slavic and German peasants from near Hungary. This is not a controversial claim on my part; it's backed up by historical sources and a wide range of genetic analyses. Hungarians still show some minor ancestry from Hungarian Conquerors (early Magyars), but this signal only reliably shows up in large surveys of Y-chromosome samples. The Scythians that you used to model the ancestry of present-day Hungarians are of local, Pannonian origin, and they don't show any eastern nomad ancestry. So they're either acculturated Scythians, or, more likely, wrongly classified as Scythians by archeologists. And since these so-called Scythians lack eastern nomad ancestry, the similarity between them and present-day Hungarians is not a sign of the impact from Avars, Hungarian Conquerors and the like, but rather a lack of significant input from such groups in present-day Hungarians.Citation... Speidel et al., High-resolution genomic ancestry reveals mobility in early medieval Europe, bioRxiv, Posted March 19, 2024, doi: https://doi.org/10.1101/2024.03.15.585102 See also... Wielbark Goths were overwhelmingly of Scandinavian origin
Friday, November 10, 2023
Wielbark Goths were overwhelmingly of Scandinavian origin
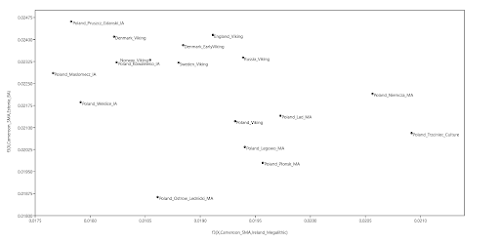
- the Wielbark Goths (Poland_Wielbark_IA) and Medieval Poles (Poland_Middle_Ages) are two distinct populations - moreover, the Wielbark Goths form a relatively compact Scandinavian-related cluster and must surely represent a homogenous population overwhelmingly of Scandinavian origin - on the other hand, the Medieval Poles form a more extensive and heterogeneous cluster that overlaps with present-day groups all the way from Central Europe to the East Baltic, and that's because they are likely to be in large part of mixed origin - I know for a fact that at least some of these early Poles harbor recent admixture, because their burials are similar to those of Vikings and their haplotypes have been shown to be partly of Scandinavian origin (see here) - one of the Wielbark females is an obvious genetic outlier (Poland_Wielbark_IA_outlier), and basically looks like a first generation mixture between a Goth and a Balt.Please note that the PCA is only based on relatively high quality genomes, so as not to confuse the picture with spurious results and noise. Also, all outliers with potentially significant ancestry from outside of Central, Eastern and Northern Europe were removed from the analysis. The relevant datasheet is available here. However, sanity checks are always important when studying complex topics like fine-scale genetic ancestry. To that end I've prepared a graph based on f3-statistics of the form f3(X,Cameroon_SMA,Estonia_BA)/(X,Cameroon_SMA,Ireland_Megalithic), that reproduces the key features of my PCA. The relevant datasheet is available here. Polish groups from the Middle Ages are marked with the MA suffix, while the Iron Age Wielbark Goths are marked with the IA suffix. If you're wondering why I plotted the f3-statistics that I did, take a look at this (all groups largely of Scandinavian origin are emboldened):
f3(X,Estonia_BA,Cameroon_SMA) Poland_Legowo_MA 0.226406 Poland_Ostrow_Lednicki_MA 0.225996 Poland_Plonsk_MA 0.225017 Poland_Trzciniec_Culture 0.224215 Poland_Lad_MA 0.224142 Poland_Viking 0.223838 Poland_Niemcza_MA 0.223659 Poland_Weklice_IA 0.223549 Poland_Kowalewko_IA 0.222584 Poland_Pruszcz_Gdanski_IA 0.222324 Sweden_Viking 0.222091 Russia_Viking 0.222042 Poland_Maslomecz_IA 0.221914 Norway_Viking 0.221825 Denmark_EarlyViking 0.221257 Denmark_Viking 0.221174 England_Viking 0.220979 f3(X,Ireland_Megalithic,Cameroon_SMA) Poland_Maslomecz_IA 0.219816 Poland_Weklice_IA 0.219501 Denmark_Viking 0.2192 Poland_Kowalewko_IA 0.219176 Poland_Ostrow_Lednicki_MA 0.218916 Norway_Viking 0.218854 Poland_Pruszcz_Gdanski_IA 0.218684 Sweden_Viking 0.218626 Denmark_EarlyViking 0.218529 England_Viking 0.218308 Russia_Viking 0.217999 Poland_Viking 0.217914 Poland_Plonsk_MA 0.217756 Poland_Lad_MA 0.217719 Poland_Legowo_MA 0.21765 Poland_Niemcza_MA 0.217001 Poland_Trzciniec_Culture 0.216551Interestingly, the Middle Bronze Age samples associated with the Trzciniec Culture (Poland_Trzciniec_Culture) show a closer genetic relationship to Medieval Poles than to Wielbark Goths or Northwestern Europeans. This is indeed the case both in terms of genome-wide and uniparental markers, including some very specific lineages under Y-chromosome haplogroup R1a. But that's a much more complex issue that I'll leave for another time. So please stay tuned. See also... Slavs have little, if any, Scytho-Sarmatian ancestry
Tuesday, June 21, 2022
My take on the Erfurt Jews
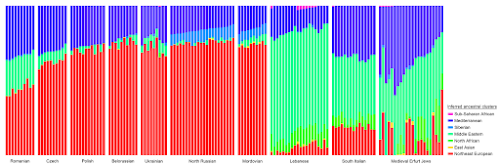
- in terms of fine scale ancestry, the Erfurt Jews show enough variation to be divided into three or four clusters, as opposed to just two as per Waldman et al. - some of the Erfurt Jews show excess "Mediterranean" ancestry, while others excess "North African" ancestry, and this cannot be explained with ancestral populations similar to Lebanese and/or South Italians, but rather with significant gene flow from the western Mediterranean and possibly North Africa - several of the Erfurt Jews show relatively high levels of "East Asian" ancestry that cannot be explained by admixture from Russians, or even any Russian-like populations, because such populations almost lack this type of ancestry, and instead show significant "Siberian" admixture - as far as I can see, there are no correlations between any of the observations above and the quality of the samples. That is, low coverage doesn't appear to be causing the aforementioned excess "Mediterranean", "North African" and/or "East Asian" ancestry proportions.Investigating this in more detail with, say, formal statistics will take some time. But I was able to reproduce the results from the above ADMIXTURE run using several somewhat different datasets, so that's something. It seems to me that Waldman et al. want a simple and elegant model to explain the data, which is understandable, but I do think they should at least expand their ADMIXTURE analysis to include "Siberian", "Mediterranean" and "North African" clusters, and go from there depending on what they find. Citation... Waldman et al., Genome-wide data from medieval German Jews show that the Ashkenazi founder event pre-dated the 14th century, bioRxiv, posted May 16, 2022, doi: https://doi.org/10.1101/2022.05.13.491805 See also... Mediterranean PCA update
Saturday, March 12, 2022
Lousy intel
Sunday, January 23, 2022
Para-Turbo-Balto-Slavic?
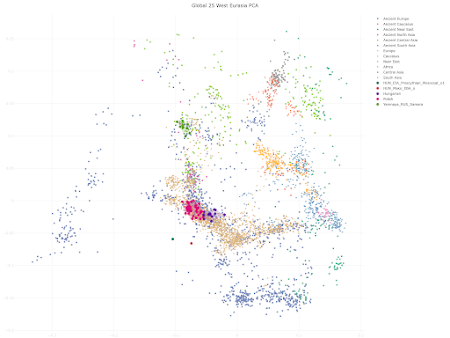
- shared genetic drift with present-day Balto-Slavic speakers to the exclusion of most other Europeans - and yet, an unusually low level of Yamnaya-related steppe ancestry - so much so, in fact, that they're often outside the range of modern European genetic variation.As far as I can tell, currently the best examples of this unusual population are HUN_Mako_EBA_o:I1502 (Mathieson et al. Nature 2015) and HUN_EIA_Prescythian_Mezocsat_o1:I18241 (Patterson et al. Nature 2021). Both are from the Carpathian Basin in what is now Hungary. I ran a series of qpAdm mixture models to try and learn more about their origins. The most robust outcomes, out of about 50 different attempts, are these:
right pops: CMR_Shum_Laka_8000BP MAR_Taforalt IRN_Ganj_Dareh_N Levant_PPNB TUR_Barcin_N Iberia_Southeast_Meso UKR_Meso England_Meso RUS_Karelia_HG RUS_West_Siberia_HG MNG_North_N TWN_Hanben BRA_LapaDoSanto_9600BP HUN_Mako_EBA_o Baltic_LTU_Narva 0.149 ∓0.028 POL_Globular_Amphora 0.613 ∓0.028 Yamnaya_RUS_Samara 0.238 ∓0.029 chisq 10.836 tail prob 0.370463 Full output HUN_EIA_Prescythian_Mezocsat_o1 Baltic_LTU_Narva 0.186 ∓0.028 POL_Globular_Amphora 0.592 ∓0.027 Yamnaya_RUS_Samara 0.222 ∓0.029 chisq 12.492 tail prob 0.253499 Full outputCombining the two genomes produces a very similar result:
HUN_EBA-EIA_o Baltic_LTU_Narva 0.160 ∓0.023 POL_Globular_Amphora 0.612 ∓0.023 Yamnaya_RUS_Samara 0.227 ∓0.023 chisq 14.653 tail prob 0.14524 Full outputImportantly, when I move RUS_Karelia_HG from the right pops to the left pops, to test whether HUN_EBA-EIA_o really has steppe ancestry, as opposed to closely related hunter-gatherer ancestry, I still get a very similar outcome:
HUN_EBA-EIA_o Baltic_LTU_Narva 0.158 ∓0.027 POL_Globular_Amphora 0.605 ∓0.033 RUS_Karelia_HG 0.014 ∓0.038 Yamnaya_RUS_Samara 0.223 ∓0.053 chisq 10.461 tail prob 0.234171 Full outputSo these largely Globular Amphora-related individuals do harbor as much as a quarter of steppe ancestry, which is to be expected considering the massive genetic turn-over that most of Europe experienced just before their time as a result of population expansions from the Pontic-Caspian steppe. Nevertheless, this is ~20% less steppe ancestry than in the present-day populations of the region, and it clearly shows in any decent Principal Component Analysis (PCA) of West Eurasia. For instance: At the same time, the relatively close genetic relationship between these ancients and present-day Balto-Slavic speaking populations shows up in fine-scale intra-European PCA. The origins and implications of this population are still a mystery to me. I don't think it's native to the Carpathian Basin. Indeed, my qpAdm models suggest that it may have moved into this region from somewhere to the northeast, because its ancestry is best modeled with ancient groups from present-day Lithuania, Poland and Russia. I'm adamant that these people weren't Balto-Slavic speakers, and certainly not proto-Slavs. Rather, I suspect that much like the Welzin warriors of Bronze Age North-Central Europe, they were closely related to a contemporaneous group that eventually gave rise to proto-Slavs. At best, they may have somehow contributed to the ethnogenesis of Balto-Slavs. By the way, using the Global25 to model their ancestry is highly problematic, because of the strong Balto-Slavic genetic drift that affects some of the dimensions. So be careful when you try it, or better yet, don't try it at all, and stick to formal stats in this particular instance. See also... Tollense Valley Bronze Age warriors were very close relatives of modern-day Slavs
Tuesday, January 11, 2022
Population genetics is a state of mind
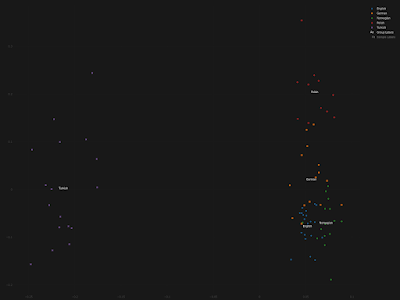
Chimp Yamnaya_Samara Swedish_modern Polish_modern -0.000311 -1.574 Chimp Yamnaya_Samara Ollsjo_Bronze_Age Polish_modern -0.000044 -0.152 Chimp Yamnaya_Samara Sealand_Iron_Age Polish_modern -0.000072 -0.293 Chimp Yamnaya_Samara Sealand_Viking_Age Polish_modern 0.000078 0.525 Chimp Yamnaya_Samara Gotland_Viking_Age Polish_modern -0.000141 -1.322 Chimp Barcin_N Swedish_modern Polish_modern -0.000318 -1.662 Chimp Barcin_N Ollsjo_Bronze_Age Polish_modern 0.000216 0.798 Chimp Barcin_N Sealand_Iron_Age Polish_modern -0.000023 -0.104 Chimp Barcin_N Sealand_Viking_Age Polish_modern -0.000186 -1.310 Chimp Barcin_N Gotland_Viking_Age Polish_modern 0.000083 0.788 Chimp Karelia_HG Swedish_modern Polish_modern -0.000134 -0.540 Chimp Karelia_HG Ollsjo_Bronze_Age Polish_modern 0.000056 0.162 Chimp Karelia_HG Sealand_Iron_Age Polish_modern 0.000047 0.153 Chimp Karelia_HG Sealand_Viking_Age Polish_modern 0.000424 2.241 Chimp Karelia_HG Gotland_Viking_Age Polish_modern 0.000134 0.959Simply put, if Poles have ~25% ancestry from a Turkish-related source, then so do Swedes, Norwegians and basically all other Northern Europeans going back hundreds and even thousands of years. This is obviously not the case, and it's also not what Hellenthal et al. claimed anyway. A year later, a team of scientists that again included Garrett Hellenthal, George B. J. Busby, James F. Wilson, Cristian Capelli and Simon Myers, published another, similar paper in Current Biology. And guess what? This paper also claimed that present-day Poles had Turkish-related ancestry, but this time dating to a somewhat later period. See Busby et al. 2015 Figure 4.C here. I've got most of the samples from that paper, so I can analyze them myself, and I think I know what the problem is. Basically, the Turks are mixed. So what appears to have happened is that Busby et al. got things backwards. Below are three plots from a Principal Component Analysis (PCA) largely based on data from Busby et al., featuring samples from England, Germany, Norway, Poland and Turkey. The first plot is based on dimensions 1 and 2, the second plot on dimensions 1 and 3, and the third plot on dimensions 1 and 4. The relevant data file is available here. Note that the Europeans are more or less symmetrically related to the Turks, which means none of these European populations has significantly more Turkish-related ancestry than the others. Indeed, it's the Turks who show more variation in the first (horizontal) dimension, suggesting that they might have variable levels of European ancestry. I chose the aforementioned papers to make my point here because they made quite an impression on me. In other words, they really pissed me off. For the sake of completeness, I'm now going to try and get in touch with the authors and ask them how on earth they managed to make these Poles Turkish-related, and also why they never corrected their mistake. See also... Don't believe everything you read in peer reviewed papers
Friday, November 13, 2020
Fatyanovo as part of the wider Corded Ware family (Nordqvist and Heyd 2020)
In the traditional narrative, the Fatyanovo people – like the CWC populations in general – are regarded as Indo-European, representing the pre-Balto-Slavic (-Germanic) stage (Carpelan & Parpola 2001, 88; Anthony 2007, 380; also Gimbutas 1956, 163; Tretyakov 1966, 109) in the spread of Indo-European languages.That's correct, but considering the latest ancient DNA research on the Fatyanovo people, the traditional narrative is probably wrong. Fatyanovo males were rich in Y-haplogroup R1a-Z93, which is found at very low frequencies in Balto-Slavic populations (see here). It's actually much more common nowadays in Central and South Asia, where it often reaches frequencies of over 50% in Indo-Iranian speaking groups. Balts and Slavs are rich in R1a-Z282, which is a sister clade of R1a-Z93 that has been found in Corded Ware and Corded Ware-related samples from west of Fatyanovo sites. That is, in present-day Poland and the Baltic states. Therefore, the origins of the Balto-Slavs should be sought somewhere west of the Fatyanovo culture, probably in the Corded Ware derived populations from what is now the border zone between Poland, Belarus and Ukraine. Indeed, in my view the Fatyanovo people are more likely to have spoken Proto-Indo-Iranian rather than anything ancestral to Baltic or Slavic (see here). Nordqvist and Heyd, The Forgotten Child of the Wider Corded Ware Family: Russian Fatyanovo Culture in Context, Proceedings of the Prehistoric Society, online 12 November 2020, DOI: https://doi.org/10.1017/ppr.2020.9 See also... The oldest R1a to date
Tuesday, September 29, 2020
Viking world open analysis and discussion thread
Tuesday, September 8, 2020
Warriors from at least two different populations fought in the Tollense Valley battle
- this may have been a battle between two Central European armies, one rich in Y-haplogroup R1b and the other rich in Y-haplogroup I2a, as well as their allies or hired help, including warriors from Eastern Europe belonging to Y-haplogroup R1a - or perhaps it was an invasion from the east by warriors rich in Y-haplogroup R1a, and it was a success, with the local armies, rich in Y-haplogroups R1b and I2a, losing the battle and suffering most of the casualties.I'm sure that one day someone will attempt to undertake a decent multidisciplinary study of this epic battle, and we'll at least have a rough idea about what happened. Or not. Citation... Burger et al., Low Prevalence of Lactase Persistence in Bronze Age Europe Indicates Ongoing Strong Selection over the Last 3,000 Years, Current Biology, Available online 3 September 2020, https://doi.org/10.1016/j.cub.2020.08.033 See also... Genetic and linguistic structure across space and time in Northern Europe
Wednesday, August 19, 2020
Yamnaya-related ancestry proportions in present-day Poles
right pops:CMR_Shum_Laka_8000BP
MAR_Taforalt
IRN_Ganj_Dareh_N
Levant_PPNB
GEO_CHG
TUR_Barcin_N
RUS_Piedmont_En
SRB_Iron_Gates_HG
WHG
RUS_Karelia_HG
MNG_North_N
RUS_Ust_Kyakhtaleft pops:Polish_East
CWC_Baltic_early 0.572±0.024
SWE_TRB 0.428±0.024
chisq 11.776
tail prob 0.300296
Full outputPolish_West
CWC_Baltic_early 0.587±0.021
SWE_TRB 0.413±0.021
chisq 11.165
tail prob 0.34478
Full output
Even using transversion sites, this is one of the very few combinations of ancient reference samples that works for the Poles with these right pops. That is, the combination of early Corded Ware samples from the East Baltic (CWC_Baltic_early) and Funnel Beaker samples from Scandinavia (SWE_TRB). The former are obviously the proxy here for Yamnaya-related ancestry. Adding any sort of hunter-gatherer population to this model doesn't help or even makes things worse (for instance, see here and here). It is possible to add Baltic hunter-gatherers to a similar model after dropping CWC_Baltic_early in favor of closely related samples from the Early to Middle Bronze Age Pontic-Caspian steppe. Note, however, that the statistical fits are somewhat poorer.
Interestingly, but not surprisingly, the ancestry of many present-day Northwestern European populations can be modeled in basically the same way. That's because ancient ancestry proportions are more closely correlated with latitude than longitude across much of the European continent.Polish_East
Baltic_LTU_Narva 0.032±0.014
PC_steppe_EMBA 0.483±0.019
SWE_TRB 0.485±0.019
chisq 17.143
tail prob 0.0465198
Full outputPolish_West
Baltic_LTU_Narva 0.031±0.011
PC_steppe_EMBA 0.491±0.015
SWE_TRB 0.477±0.016
chisq 22.444
tail prob 0.00757421
Full output
A zip file with the qpAdm output from this analysis and a list of the most relevant ancients is available here. I might try to run a few more populations over the next few days, but probably only from the northern half of Europe, so please check the zip file in a week or so to see what else is in there. If anyone wants to challenge my results, note that these and very similar samples are freely available to the public via Harvard University here and here. Update 22/08/2020: From Nick Patterson (Broad) in the comments:English_Kent
CWC_Baltic_early 0.527±0.024
SWE_TRB 0.473±0.024
chisq 13.042
tail prob 0.221357
Full outputIcelandic
CWC_Baltic_early 0.586±0.023
SWE_TRB 0.414±0.023
chisq 16.517
tail prob 0.085751
Full outputScottish
CWC_Baltic_early 0.583±0.021
SWE_TRB 0.417±0.021
chisq 12.144
tail prob 0.275536
Full output
My general advice for qpAdm is 1) Work on the right hand set. Don't include irrelevant population (except for one population as an outgroup); picking the best RHS can dramatically reduce s. errors on the admixture weights. 2) If qpAdm gives a very low p-value try and understand why, sometimes it is telling you that the target is not a mixture of the sources but sometimes the assumptions are violated, for example recent gene-flow from left pops -> right.See also... Ancient ancestry proportions in present-day Europeans
Monday, July 27, 2020
Ancient ancestry proportions in present-day Europeans (to be continued)
TUR_Barcin_N
UKR_Yamnaya
MAR_Taforalt
Levant_Natufian
IRN_Ganj_Dareh_N
Levant_PPNB
CZE_Vestonice16
BEL_GoyetQ116-1
Iberia_ElMiron
RUS_Karelia_HG
RUS_West_Siberia_HG
MNG_North_N
RUS_Ust_Kyakhta
HUN_Koros_N_HG 0.134±0.043
RUS_Baikal_BA 0.270±0.015
TUR_Barcin_N 0.081±0.026
UKR_Yamnaya 0.515±0.058
Monday, July 13, 2020
Don't believe everything you read in peer reviewed papers
The Mordovian and Csango samples have a moderate to slight orientation toward the Central-Asian and Siberian Turkic groups. This could suggest the more significant East Eurasian or Turkic ancestry of these populations, which should be further investigated. German samples are inhomogeneous, and some of the German samples also show this tendency, which can be the result of the recent 20th century Turkish immigration into Germany [42].Nope, these German samples don't show anything even remotely resembling recent Turkish ancestry. The authors of the paper, Ádám, V., Bánfai, Z., Maász, A. et al., should've been able to figure this out, even with the standard analyses that they ran. Failing that, the peer reviewers at the Journal of Human Genetics should've noticed that the authors were confused. Moreover, if the authors and peer reviewers actually bothered to take a closer look at metadata for these samples, which were sourced from the Estonian Biocentre, they'd see that they're not even from Germany. In fact, they represent self-reported ethnic Germans from Russia. My own quick and dirty analysis of these individuals suggests that many of them harbor East Slavic and/or Volga Finnic ancestries. Indeed, only some of them can pass genetically for run of the mill Germans from Germany. The Principal Component Analysis (PCA) below is self-explanatory. It was plotted with the Vahaduo Custom PCA tools freely available here. The relevant PCA datasheet can be gotten here. That's not to say, of course, that some Germans don't have recent Turkish ancestry, because an increasing number of Germans nowadays do, nor that people with German heritage in Russia shouldn't identify as Germans, because that's entirely their choice. This blog post isn't about what it takes to be German, and this is not something that I ever want to discuss for obvious reasons. The point I'm making here is that the authors and peer reviewers of the said paper at the Journal of Human Genetics were sloppy and half-arsed in their approach. And, sadly, this isn't an isolated case in peer reviewed scientific literature dealing with human population genetics. I feel that the Estonian Biocentre is also partly to blame for this cock up, due to its somewhat peculiar sampling and labelling strategies. For instance, its scientists rely solely on self-reported identity to establish the ethnic origins of their samples, and they apparently never remove genetic outliers from their datasets or even try to identify them. Unfortunately, I fear that this relaxed approach will eventually lead to basic errors and even unusual conclusions in a number of so called peer reviewed papers. I first raised this issue with the Estonian Biocentre about five years ago, when I noticed that some of the supposedly Polish individuals in its dataset were genetically more similar to various groups from northern Russia than to Poles from Poland. These individuals also showed significant Siberian ancestry, which was very unusual indeed. Where the hell did the Estonian Biocentre find Poles who resembled people from near the Arctic Circle, you might ask? Apparently in Estonia. OK, I can imagine that sampling ethnic Poles from Estonia may have been easier for the Estonian Biocentre than sampling Poles from Poland. And Estonian Poles certainly make for interesting and useful data points. However, as you can see in the PCA below, some of these individuals (labeled Polish_Estonia by me) aren't representative of the native Polish population, and yet the Estonian Biocentre not only lumps them with their Poles from Poland, but even labels them with the word "Poland". The relevant PCA datasheet can be gotten here. However, based on my communications with some of the scientists at the Estonian Biocentre, including head honcho Mait Mestpalu, it seems that nothing will ever change there in regards to this issue. Who knows, perhaps some day we'll see a paper based on Estonian Biocentre data in the Journal of Human Genetics claiming that Poles originated near the Arctic Circle? I wouldn't be shocked if that actually happened. Citation... Ádám, V., Bánfai, Z., Maász, A. et al. Investigating the genetic characteristics of the Csangos, a traditionally Hungarian speaking ethnic group residing in Romania. J Hum Genet (2020). https://doi.org/10.1038/s10038-020-0799-6 See also... Like three peas in a pod
Friday, April 17, 2020
Corded Ware cultural and genetic complexity (Linderholm et al. 2020)
During the Final Eneolithic the Corded Ware Complex (CWC) emerges, chiefly identified by its specific burial rites. This complex spanned most of central Europe and exhibits demographic and cultural associations to the Yamnaya culture. To study the genetic structure and kin relations in CWC communities, we sequenced the genomes of 19 individuals located in the heartland of the CWC complex region, south-eastern Poland. Whole genome sequence and strontium isotope data allowed us to investigate genetic ancestry, admixture, kinship and mobility. The analysis showed a unique pattern, not detected in other parts of Poland; maternally the individuals are linked to earlier Neolithic lineages, whereas on the paternal side a Steppe ancestry is clearly visible. We identified three cases of kinship. Of these two were between individuals buried in double graves. Interestingly, we identified kinship between a local and a non-local individual thus discovering a novel, previously unknown burial custom. ... The PCA revealed that despite geographical proximity there is a distinct genetic separation between CWC and BBC individuals from southern Poland. The genetic variation of CWC individuals from southern Poland overlaps with the majority of previously published CWC individuals from Germany while the eight published CWC individuals from the Polish lowland [10,11] more closely resemble BBC individuals (Fig. S21). This fact is not unexpected if we consider the CWC communities in Polish lowlands as representatives of north-western parts of the CWC world called as the Single-Grave culture (see supplementary information). The genetic variation of BBC individuals from south-eastern Poland overlaps with the broad variation of BBC individuals from Central Europe (Bohemia, Moravia, Germany, south-western Poland and Hungary) (Fig. S22) which corresponds well with archaeological data.Linderholm, A., Kılınç, G.M., Szczepanek, A. et al. Corded Ware cultural complexity uncovered using genomic and isotopic analysis from south-eastern Poland. Sci Rep 10, 6885 (2020). https://doi.org/10.1038/s41598-020-63138-w See also... The Battle Axe people came from the steppe Is Yamnaya overrated? Single Grave > Bell Beakers
Wednesday, July 17, 2019
Viking invasion at bioRxiv
The Viking maritime expansion from Scandinavia (Denmark, Norway, and Sweden) marks one of the swiftest and most far-flung cultural transformations in global history. During this time (c. 750 to 1050 CE), the Vikings reached most of western Eurasia, Greenland, and North America, and left a cultural legacy that persists till today. To understand the genetic structure and influence of the Viking expansion, we sequenced the genomes of 442 ancient humans from across Europe and Greenland ranging from the Bronze Age (c. 2400 BC) to the early Modern period (c. 1600 CE), with particular emphasis on the Viking Age. We find that the period preceding the Viking Age was accompanied by foreign gene flow into Scandinavia from the south and east: spreading from Denmark and eastern Sweden to the rest of Scandinavia. Despite the close linguistic similarities of modern Scandinavian languages, we observe genetic structure within Scandinavia, suggesting that regional population differences were already present 1,000 years ago. We find evidence for a majority of Danish Viking presence in England, Swedish Viking presence in the Baltic, and Norwegian Viking presence in Ireland, Iceland, and Greenland. Additionally, we see substantial foreign European ancestry entering Scandinavia during the Viking Age. We also find that several of the members of the only archaeologically well-attested Viking expedition were close family members. By comparing Viking Scandinavian genomes with present-day Scandinavian genomes, we find that pigmentation-associated loci have undergone strong population differentiation during the last millennia. Finally, we are able to trace the allele frequency dynamics of positively selected loci with unprecedented detail, including the lactase persistence allele and various alleles associated with the immune response. We conclude that the Viking diaspora was characterized by substantial foreign engagement: distinct Viking populations influenced the genomic makeup of different regions of Europe, while Scandinavia also experienced increased contact with the rest of the continent.Margaryan et al., Population genomics of the Viking world, bioRxiv, posted July 17, 2019, doi: https://doi.org/10.1101/703405 See also... They came, they saw, and they mixed Who were the people of the Nordic Bronze Age? Asiatic East Germanics
Tuesday, May 7, 2019
The execution
Wednesday, December 31, 2008
Best of 2008: Corded Ware DNA from Germany
One of the biggest hits of the year for this blogger was the discovery of Y-DNA haplogroup R1a among three Corded Ware skeletons from a burial site in Eulau, eastern Germany. It's an important result, because it links one of Europe's most dominant Y-haplogroups to a major Late Neolithic archeological complex.
All three individuals were confirmed to be paternally related via their shared Y-STR haplotype. Nevertheless, the outcome appears far from a random coincidence. Consider that in Europe today R1a shows its highest frequencies in Poland and Western Russia, which are both located in former Corded Ware territory, and where the Eulau R1a haplotype appears to have its closest modern matches. Moreover, the Corded Ware culture is often classified as an Indo-European culture by archeologists and linguists, while at the same time R1a has been posited as a marker of the early Indo-Europeans by some geneticists. Needless to say, I'm expecting R1a to be a common, and perhaps dominant marker among Corded Ware samples when more of them make it to the lab.
The consensus haplotype of the three individuals (based on most complete profile) gave two exact matches in in an European population sample of 11,213 haplotypes in a set of 100 populations (as of July 2008, Release ‘‘23’’ from 2008–01-15 14:44:25): one individual from Poland (1/939 from Gdansk) and one from Russia (1/48 from Tambov).
...
The Y haplotype was predicted using the Web-based program Haplotype Predictor (9). The three individuals of grave 99 belong to haplotype R1a, with a probability of 100% based on the Y-STR profile of individual 3 (10). To confirm haplogroup status, we further amplified an 85-bp fragment covering the Y-SNP marker SRY10831.2 characteristic for R1a (11). Primer sequences are given in Table S6. Sequences and sequenced clones from independent extract of all three individuals show the specific G to A transition identifying R1a (Fig. S5).
The mitochondrial DNA (mtDNA) lineages of the Eulau skeletons belonged to haplogroups K1b (3), X2 (2), H, I, K1a2, and U5b. Most of these maternal markers aren't particularly common in Europe today, and the overall result appears decidedly unusual compared to the mtDNA frequencies of modern European populations, largely because of the low frequency of H.
I'm quite certain this is at least partly due to the small sample size and presence of several related individuals skewing some of the frequencies. However, it's interesting to note that this pattern of discontinuity between mtDNA gene pools from different time periods has also been reported in other studies, some with larger samples, and focusing on different regions of Europe. So it might well be a signal of significant shifts in mtDNA frequencies during European prehistory and early history, possibly as a result of major migrations leading to significant population replacements.
Interestingly, one of the ancient K1b lineages most closely matched a haplotype shared by two modern Shugnans from Tajikistan. Exactly how the Corded Ware individual is related to these two Central Asians isn't clear yet, but Shugni is an Indo-Iranian language, so some kind of early Indo-European relationship is possible.
Citation...
Wolfgang Haak et al, Ancient DNA, Strontium isotopes, and osteological analyses shed light on social and kinship organization of the Later Stone Age, PNAS, Published online before print November 17, 2008, doi:10.1073/pnas.0807592105