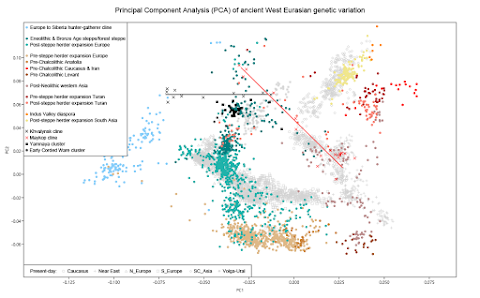
This direct evidence that most Corded Ware ancestry must have genealogical links to people associated with Yamnaya culture spanning on the order of at most a few hundred years is inconsistent with the hypothesis that the Steppe-like ancestry in the Corded Ware primarily reflects an origin in as-of-now unsampled cultures genetically similar to the Yamnaya but related to them only a millennium earlier.This is basically a straw man argument, because it's easy to debunk. So why put it in the paper? Well, as far as I can see, to make the idea that the CWC is derived from Yamnaya look more plausible. This idea, that the CWC is an offshoot of Yamnaya, seems to be the favorite explanation for the appearance of the CWC among the scientists at the David Reich Lab. However, I'd say they're facing a major problem with that, because the CWC and Yamnaya populations have largely different paternal origins. That is, CWC males mostly belong to Y-haplogroups R1a-M417 and R1b-L51, while Yamnaya males almost exclusively belong to Y-haplogroup R1b-Z2103. Indeed, as far as I know, there are no reliable instances of R1a-M417 or R1b-L51 in any published or yet to be published Yamnaya samples. But it is possible to reconcile the Y-haplogroup data with the ancIBD results if we assume that the peoples associated with the Corded Ware, Yamnaya and also Afanasievo cultures expanded from a genetically more diverse ancestral gene pool, each taking a specific subset of the variation with them. This gene pool would've existed somewhere in Eastern Europe, probably at the western end of the Pontic-Caspian steppe, at most a few hundred years before the appearance of the earliest CWC burials in what is now Poland. Moreover, the split between the CWC and Yamnaya populations need not have been a clean one, with long-range contacts and largely female-mediated mixing maintained for generations, adding to the already close genealogical links between them. Citation... Ringbauer, H., Huang, Y., Akbari, A. et al. Accurate detection of identity-by-descent segments in human ancient DNA. Nat Genet (2023). https://doi.org/10.1038/s41588-023-01582-w See also... On the origin of the Corded Ware people
search this blog
Showing posts with label CWC. Show all posts
Showing posts with label CWC. Show all posts
Wednesday, December 20, 2023
Dear Harald #2
The ancIBD method paper from the David Reich Lab was just published in Nature (open access here). It's a very useful effort, but the authors are still somewhat confused about the origin of the Corded Ware culture (CWC) population. From the paper (emphasis is mine):
Saturday, April 8, 2023
Dear Harald...
I've started analyzing the Identity-by-Descent (IBD) data from the recent Ringbauer et al. preprint (see here). Unfortunately, it'll take me a few weeks to do this properly, so I won't be able to write anything detailed on the topic for a while.
Meantime, this is the comment that I left for the authors at bioRxiv (at this time it's still being approved, but it should appear there within a day or so, possibly along with a reply from the authors):
Hello authors, Thanks for the interesting preprint and data. However, I'd like to see you address a couple of technical issues and perhaps one theoretical issue in the final manuscript: - the output you posted shows some unusual results, which are potentially false positives that appear to be concentrated among the shotgun and noUDG samples. I'm guessing that this is due to the same types of ancient DNA damage creating IBD-like patterns in these samples. If so, isn't there a risk that many or even most of the individuals in your analysis are affected by this problem to some degree, which might be skewing your estimates of genealogical relatedness between them? - many individuals from groups that have experienced founder effects, such as Ashkenazi Jews, appear to be close genetic cousins, even though they're not genealogical cousins. Basically, the reason for this is reduced haplotype diversity in such populations. Have you considered the possibility that at least some of the close relationships that you're seeing between individuals and populations might be exaggerated by founder effects? - thanks to ancient DNA we've learned that the Yamnaya phenomenon isn't just an archeological horizon, but also a closely related and genetically very similar group of people. Indeed, in my mind, ancient DNA has helped to redefine the Yamnaya concept, with Y-chromosome haplogroup R1b-Z2103 now being one of the key traits of the Yamnaya identity. So considering that the Corded Ware people are not rich in R1b-Z2103, and even the earliest Corded Ware individuals are somewhat different from the Yamnaya people in terms of genome-wide genetic structure, it doesn't seem right to keep claiming that the Corded Ware population is derived from Yamnaya. I can't see anything in your IBD data that would preclude the idea that the Corded Ware and Yamnaya peoples were different populations derived from the same as yet unsampled pre-Yamnaya/post-Sredny steppe group.See also... Dear Harald #2 On the origin of the Corded Ware people
Labels:
ancIBD,
ancient ancestry,
ancient DNA,
Corded Ware Culture,
CWC,
David Reich,
Eastern Europe,
haplotype,
Harald Ringbauer,
IBD,
Identity-by-Descent,
Nick Patterson,
Pontic-Caspian steppe,
Yamnaya
Friday, January 21, 2022
Yamnaya is from Europe, but it's really from Asia
I was about to post a comment under a new preprint at bioRxiv, but the comment section isn't there anymore. Hopefully, this is just a temporary glitch.
The preprint in question is titled Reconstructing the spatiotemporal patterns of admixture during the European Holocene using a novel genomic dating method [LINK]. It's co-authored by Harvard/Broad MIT scientist Nick Patterson who occasionally comments at this blog.
My impression is that the authors see the people associated with the Yamnaya culture as Asians who simply used "far" Eastern Europe as a springboard to expand into other parts of Europe.
If so, they're dead wrong.
There are at least three arguments why the Yamnaya population should be seen as quintessentially European:
- its home was initially and overwhelmingly the Pontic-Caspian steppe, which is entirely located within the present-day borders of Europe - Yamnaya genomes are clearly different from those of older populations native to nearby parts of Asia, and, in fact, these differences show a very strong correlation with the present-day borders between Europe and Asia - the Yamnaya people weren't a new population in Europe by any stretch, but must have been overwhelmingly derived from the very similar Eneolithic peoples of the Pontic-Caspian steppe and/or the nearby forest steppe, both of which are located in Eastern Europe.And yet, this is what the preprint claims:
The beginning of the Bronze Age was a period of major cultural and demographic change in Eurasia, accompanied by the spread of Yamnaya Steppe Pastoralist-related ancestry from Pontic-Caspian steppes into Europe and South Asia (16).In fact, what really happened at this time was that Yamnaya steppe pastoralist-related ancestry spread from Eastern Europe to other parts of Europe, as well as to Central and West Asia. The preprint does eventually explain that present-day South Asians derive their Yamnaya-related ancestry from a later eastward expansion of the European Corded Ware culture (CWC), but it completely ignores the fact that the Afanasievo culture was the result of the initial eastward expansion from Europe to Asia. That is, the ancestors of the Afanasievo people were recent migrants from the Pontic-Caspian steppe to Central Asia and Siberia. There's also this:
Over the following millennium, the Yamnaya-derived groups of the Corded Ware Complex (CWC) and Bell Beaker complex (BBC) cultures brought Steppe pastoralist-related ancestry to Europe.Seriously? Both the CWC and BBC, just like the Yamnaya culture, were from Europe. In fact, as per above, the descendants of the CWC expanded into Asia. And this:
The second major migration occurred when populations associated with the Yamnaya culture in the Pontic-Caspian steppe expanded to central and western Europe from far eastern Europe.The authors basically admit here that Yamnaya came from Eastern Europe, but they call it "far" Eastern Europe. Perhaps they know something I don't, but as things stand, there's no evidence that Yamnaya came from "far" Eastern Europe. In fact, the emerging consensus based on ancient DNA, including pre-publication data, is that Yamnaya may have originated in what is now Ukraine. In my opinion, Ukraine isn't located in "far" Eastern Europe, but more or less in the middle of it. Inexplicably, this is what they say about the genetic origins of the Yamnaya and Afanasievo peoples:
These groups were likely the result of a genetic admixture between the descendants of EHG-related groups and CHG-related groups associated with the first farmers from Iran (8, 22, 36). ... Thus, we combined all early Steppe pastoralist individuals in one group to obtain a more precise estimate for the genetic formation of proto-Yamnaya of ~4,400 to 4,000 BCE (Figure 2). These dates are noteworthy as they pre-date the archeological evidence by more than a millennium (37) and have important implications for understanding the origin of proto-Pontic Caspian cultures and their spread to Europe and South Asia.Not really. Like I said, the Yamnaya population was overwhelmingly derived from the Eneolithic peoples of the Eastern European steppe and/or forest steppe. And these Yamnaya-like Eneolithic peoples were spread out across a vast area of Eastern Europe by at least ~4,500 BCE. Some of their genomes have been available for several years, and many more are on the way. It is possible that the Yamnaya and Afanasievo genotype formed in 4,400-4,000 BCE, but if so, then this was due to mixing between the Eneolithic steppe peoples and nearby European farmers. That's because the difference between the Yamnaya and Eneolithic steppe genotypes is minor (~15%) European farmer admixture in the former. The really interesting puzzle is exactly where and when the peculiar Eneolithic steppe genotype came into being. Any ideas Dr Patterson? See also... Matters of geography Understanding the Eneolithic steppe
Labels:
Afanasevo,
Afanasievo,
ancient DNA,
Corded Ware,
Corded Ware Culture,
CWC,
Eastern Europe,
Eurasia,
Eurasian steppe,
Indo-European,
Pontic-Caspian steppe,
Proto-Indo-European,
Yamna,
Yamnaya
Friday, August 27, 2021
R1a vs R1b in third millennium BCE Central Europe (Papac et al. 2021)
R1a-M417 and R1b-L51 are by far the most important Y-chromosome haplogroups in Europe today. More precisely, R1a-M417 dominates in Eastern Europe, while R1b-L51 in Western Europe.
It's been obvious for a while now, at least to me, that both of these Y-haplogroups are closely associated with the men of the Late Neolithic Corded Ware culture (CWC). Indeed, in my mind they're the main genetic signals of its massive expansion, probably from a homeland somewhere north of the Black Sea in what is now Ukraine.
I'm still not exactly sure how the east/west dichotomy between R1a and R1b emerged in Europe, but, thanks to a new paper by Papac et al. at Science Advances, at least now I have a working hypothesis about that. Below is a quote from the said paper, emphasis is mine:
In addition to autosomal genetic changes through time, we observe a sharp reduction in Y-chromosomal diversity going from five different lineages in early CW to a dominant (single) lineage in late CW (Fig. 4A). We used forward simulations to explore the demographic scenarios that could account for the observed reduction in Y-chromosomal diversity. Performing 1 million simulations of a population with a starting frequency of R1a-M417(xZ645) centered around the observed starting frequency in Bohemia_CW_Early (3 of 11, 0.27), we assessed the plausibility of this lineage reaching the observed frequency in Bohemia_CW_Late (10 of 11, 0.91) in the time frame of 500 years under a model of a closed population and random mating (Materials and Methods). We reject the “neutral” hypothesis, i.e., that this change in frequency occurred by chance, given a wide range of plausible population sizes. Instead, our results suggest that R1a-M417(xZ645) was subject to a nonrandom increase in frequency, resulting in these males having 15.79% (4.12 to 44.42%) more surviving offspring per generation relative to males of other Y-haplogroups. We also find that this change in Y chromosome frequency is extreme compared to the changes in allele frequencies at fully covered autosomal 1240k sites within the same males, suggesting a process that disproportionately affected Y-chromosomal compared to autosomal genetic diversity, ruling out a population bottleneck as the likely cause. Our results suggest that the Y-lineage diversity in early CW males was supplanted by a nonrandom process [selection, social structure, or influx of nonlocal R1a-M417(xZ645) lineages] that drove the collapse in Y-chromosomal diversity. A simultaneous decline of Y-chromosomal diversity dating to the Neolithic has been observed across most extant Y-haplogroups (64), possibly due to increased conflict between male-mediated patrilines (65). We view that changes in social structure (e.g., an isolated mating network with strictly exclusive social norms) could be an alternative cause but would be difficult to distinguish in the underlying model parameters.Right, so even though the CWC was clearly a community of closely related groups, there must have been some competition between its different clans. And since these clans were highly patriarchal and patrilineal, this competition probably led to different paternal lineages dominating different parts of the CWC horizon, with M417 becoming especially common in the east and L51 in the west. Of course, the expansions of post-Corded Ware groups, such as the M417-rich Slavs in Eastern Europe and L51-rich Celts in Western Europe, were also instrumental in creating Europe's R1a/R1b dichotomy, but obviously these groups were in large part the heirs of the CWC. By the way, most of the samples from Papac et al. are already in the Global25 datasheets linked here. Look for the labels listed here. Below is a plot made from the Global25 data courtesy of regular commentator Matt. Citation: L. Papac, M. Ernée, M. Dobeš, M. Langová, A. B. Rohrlach, F. Aron, G. U. Neumann, M. A. Spyrou, N. Rohland, P. Velemínský, M. Kuna, H. Brzobohatá, B. Culleton, D. Daněček, A. Danielisová, M. Dobisíková, J. Hložek, D. J. Kennett, J. Klementová, M. Kostka, P. Krištuf, M. Kuchařík, J. K. Hlavová, P. Limburský, D. Malyková, L. Mattiello, M. Pecinovská, K. Petriščáková, E. Průchová, P. Stránská, L. Smejtek, J. Špaček, R. Šumberová, O. Švejcar, M. Trefný, M. Vávra, J. Kolář, V. Heyd, J. Krause, R. Pinhasi, D. Reich, S. Schiffels, W. Haak, Dynamic changes in genomic and social structures in third millennium BCE central Europe. Sci. Adv. 7, eabi6941 (2021). See also... On the origin of the Corded Ware people Understanding the Eneolithic steppe Conan the Barbarian probably belonged to Y-haplogroup R1a
Labels:
ancient DNA,
Balto-Slavic,
Celtic,
Central Europe,
Corded Ware,
Corded Ware Culture,
CWC,
Eastern Europe,
Germanic,
Indo-European,
Northern Europe,
Proto-Indo-European,
R1a-M417,
R1b-L151,
R1b-L51,
Slavic,
Slavs
Tuesday, July 20, 2021
On the origin of the Corded Ware people
There's been a lot of talk lately about the finding that the peoples associated with the Corded Ware and Yamnaya archeological cultures were genetic cousins (for instance, see here). As I've already pointed out, this is an interesting discovery, but, at this stage, it's difficult to know what it means exactly.
It might mean that the Yamnayans were the direct predecessors of the Corded Ware people. Or it might just mean that, at some point, the Corded Ware and Yamnaya populations swapped women regularly (that is, they practiced female exogamy with each other).
In any case, I feel that several important facts aren't being taken into account by most of the interested parties. These facts include, in no particular order:
- despite being closely related, the Corded Ware and Yamnaya peoples were highly adapted to very different ecological zones - temperate forests and arid steppes, respectively - and this is surely not something that happened within a few years and probably not even within a couple of generations - both the Corded Ware and Yamnaya populations expanded widely and rapidly at around the same time, but never got in each others way, probably because they occupied very different ecological niches - despite sharing the R1b Y-chromosome haplogroup, their paternal origins were quite different, with Corded Ware males rich in R1a-M417 and R1b-L51 and Yamnaya males rich in R1b-Z2103 and I2a-L699I suppose it's possible that the Corded Ware people were overwhelmingly and directly derived from the Yamnaya population. But right now my view is that, even if they were, then the Yamnaya population that they came from was quite different from the classic, R1b-Z2103-rich Yamnaya that spread rapidly across the steppes. Indeed, perhaps what we're dealing with here is a very early (proto?) Yamnaya gene pool located somewhere in the border zone between the forests and the steppes, that then split into two main sub-populations, with one of these groups heading north and the other south? I do wonder what David Anthony would say if he was made aware of the above mentioned facts? Then again, perhaps he's already aware of them, and simply chose to ignore them when formulating his latest theory about the origin of the Corded Ware people? See also...
Monday, June 28, 2021
The PIE homeland controversy: June 2021 status report
Archeologist David Anthony has made several appearances online recently to promote his theories about the origins of the Corded Ware and Yamnaya cultures and peoples.
In a clip on Youtube he reiterated his theory that the so called Iranian-related ancestry in the Yamnaya people actually came from what is now Iran, and, more precisely, that it was carried by hunter-gatherers who travelled relatively rapidly from the South Caspian region into the Volga Delta in what is now Russia.
It's still a complete mystery to me as to why a group of hunter-gatherers from the South Caspian would undertake such a migration, instead of, say, expanding their range gradually over thousands of years, first into the Caucasus and eventually into Eastern Europe.
But there's a more serious problem with Anthony's theory: it contradicts the currently available ancient DNA. That's because the so called Iranian-related ancestry in the Yamnaya people is most closely related to the Kotias and Satsurblia hunter-gatherers from what is now Georgia, and these hunter-gatherers form a separate clade from the earliest samples from what is now Iran. For instance, see here and here.
Also, in a podcast on Razib's blog, Anthony doubled down on his theory that Y-chromosome haplogroup R1a was closely associated with Yamnaya plebs who were excluded from Kurgan burials, and, as a result, their remains haven't yet been sampled.
At least this theory isn't yet contradicted by ancient DNA, but it's more complicated and less parsimonious than my theory, which posits that R1a, or rather R1a-M417, was simply a very rare lineage in the Yamnaya population, and that it only became a common and widespread marker thanks to the Corded Ware expansion (see here).
Intriguingly, my understanding is that there are several unpublished R1a samples from the Caspian and Volga steppes at Harvard's David Reich Lab that have been classified by its scientists as Yamnaya outliers. Of course, Anthony is collaborating on at least one major paper with this lab (see here).
Ergo, I strongly suspect that Anthony's theory is in part based on these Yamnaya outliers. However, I also believe that these samples are wrongly dated and probably represent Scythians and/or Sarmatians. I'll be able to look into that if they're ever published.
Speaking of the David Reich Lab, its leading scientists, David Reich and Nick Patterson, have also made appearances online recently, on Youtube and Razib's blog, respectively, to reveal that the Corded Ware and Yamnaya peoples aren't just very similar genetically, but in fact close cousins.
This is a very interesting finding. Apparently it's based on a relatively high level of Identity-by-Descent (IBD) segment sharing between Corded Ware and Yamnaya samples, but that's all I know. I'm guessing that the relevant paper is coming soon (that is, within the next five years).
However, the long-standing question that the readers of this blog want to see answered is not whether the Corded Ware and Yamnaya peoples are close cousins, but whether Yamnaya migrants founded the Corded Ware culture. The obvious way to prove that they did is to find at least one ancient population unambiguously classified as part of the Yamnaya horizon that is rich in the typically Corded Ware Y-haplogroups R1a-M417 and R1b-L151.
See also...
On the origin of the Corded Ware people
The PIE homeland controversy: January 2019 status report
The PIE homeland controversy: August 2019 status report
Labels:
ancient DNA,
Corded Ware,
CWC,
David Anthony,
David Reich,
David Reich Lab,
Eastern Europe,
Indo-European,
Iran,
Nick Patterson,
Proto-Indo-European,
R1a-M417,
R1b-L151,
R1b-L51,
Yamna,
Yamnaya
Sunday, March 31, 2019
Map of pre-Corded Ware culture (>2900 BCE) instances of Y-haplogroup R1a (updated)
Below is a map showing the global distribution of Y-chromosome haplogroup R1a prior to the expansions of the R1a-rich Corded Ware culture (CWC) people and their descendants across Europe and Asia from around 2900 BCE. I'll be updating this map regularly and using it to help me narrow down the options for the place of origin of R1a, and also to counter the misinformation about this topic that has appeared in print and online over the years, including in many scientific publications and popular websites such as Wikipedia.
Incredibly, as far as I know, there are just six reliably called instances of R1a in the now ample Eurasian ancient DNA record dating to the pre-CWC period. To put this into perspective, consider that R1a is today the most common Y-haplogroup in much of Europe and Asia. How did that happen I wonder? However, please note that I chose to base the map only on samples sequenced with the capture and shotgun methods, rather than the PCR method, which is susceptible to producing contaminated results and no longer used in major ancient DNA studies.
See also...
Y-haplogroup R1a and mental health
The Poltavka outlier
Late PIE ground zero now obvious; location of PIE homeland still uncertain, but...
Labels:
ancient DNA,
Bronze Age,
Copper Age,
Corded Ware Culture,
CWC,
Eastern Europe,
Eurasia,
PIE,
Proto-Indo-European,
R1a,
R1a origin,
R1a-M17,
R1a-M198,
R1a-M417,
R1a-M420,
R1a-Z645,
R1a-Z93,
R1a1a1,
Y-haplogroup
Tuesday, January 1, 2019
The PIE homeland controversy: January 2019 status report
Last year, the preprint that claimed to have presented archaeogenetic data that opened up the possibility of the Proto-Indo-European (PIE) homeland being located south of the Caucasus was, ironically, also the preprint that considerably strengthened my confidence that the said homeland was actually located north of the Caucasus.
Of course, I'm talking about the Wang et al. manuscript at bioRxiv, which is apparently soon to be published as a peer-reviewed paper in Nature Communications (see here).
It'll be fascinating to observe if and how the peer-review process has impacted on the preprint, and especially its conclusion. My impression was that the authors seemed pretty sure that the Maykop people gave rise to the Yamnaya culture, or at least Indo-Europeanized it. But, as far as I saw, the archaeogenetic data didn't bear this out at all, and instead showed a lack of any direct, recent and meaningful genetic relationship between Maykop and Yamnaya (see here). Was this also picked up by the peer reviewers? We shall see.
Moreover, there was some exceedingly interesting fine print in the manuscript's supplementary information:
Complementary to the southern [Darkveti-Meshoko] Eneolithic component, a northern component started to expand between 4300 and 4100 calBCE manifested in low burial mounds with inhumations densely packed in bright red ochre. Burial sites of this type, like the investigated sites of Progress and Vonyuchka, are found in the Don-Caspian steppe [10], but they are related to a much larger supra-regional network linking elites of the steppe zone between the Balkans and the Caspian Sea [16]. These groups introduced the so-called kurgan, a specific type of burial monument, which soon spread across the entire steppe zone.
Always read the fine print, they say. And they're right. Imagine if I only read the preprint's conclusion and missed this little gem; I'd probably think that the PIE homeland was located south of the Caucasus rather than on the Don-Caspian steppe.
Wow, proto-kurgans with inhumations densely packed in bright red ochre? A supra-regional network linking the elites of the steppe all way from the Balkans to the Caspian Sea? An expansionist culture? And, as evidenced by the ancient DNA from the Progress and Vonyuchka sites, a people who may well have been in large part ancestral to the Yamnaya, Corded Ware and Andronovo populations, that have been identified based on archeological and historical linguistics data as the main vectors for the spread of Indo-European languages as far as Iberia in the west and the Indian subcontinent in the east.
I wonder if the authors actually asked themselves who these people may have been, before so haphazardly turning to Maykop and, ultimately, the Near East, as the likely sources of the Yamnaya culture? To me they look like the Proto-Indo-Europeans and true antecedents of Yamnaya.
So as things stand, my pick for the PIE homeland is firmly the Don-Caspian steppe. And I genuinely thank Wang et al., and indeed the Max-Planck-Institut für Menschheitsgeschichte (aka MPI-SHH), for their assistance.
But, you might ask, what about the Hittites? Yes, I realize that no one apart from me and a few of my readers here can find any steppe ancestry in the so called Hittite genomes published to date. However, consider this: if the PIE homeland really was on the steppe, and a dense sampling strategy of Hittite era Anatolia fails to turn up any unambiguous steppe ancestry in at least a few individuals, then there has to be an explanation for it. But let's wait and see what a dense sampling strategy of Hittite era Anatolia actually reveals before we go that far.
See also...
The PIE homeland controversy: August 2019 status report
Yamnaya: home-grown
Late PIE ground zero now obvious; location of PIE homeland still uncertain, but...
Labels:
ancient DNA,
Bell Beaker,
Caucasus,
Corded Ware Culture,
CWC,
Eneolithic steppe,
Late Proto-Indo-European,
Maykop,
MPI-SHH,
mtDNA,
PIE,
Pontic-Caspian steppe,
Proto-Indo-European,
Yamnaya
Monday, December 3, 2018
On the trail of the Proto-Uralic speakers (work in progress)
Historical linguists have long posited that Fennoscandia was a busy contact zone between early Germanic and Uralic languages. The first ancient DNA samples from what is now Finland have corroborated their inferences, by showing that during the Iron Age the western part of the country was inhabited by a genetically heterogeneous population closely related to both the Uralic-speaking Saami and Germanic-speaking southern Scandinavians.
The samples were sequenced and analyzed by two different teams of researches, and their findings published recently in Lamnidis et al. and Sikora et al. (see here and here, respectively).
This is how most of these ancients, whose remains were excavated from the Levanluhta burial site dated to 300–800 CE, behave in a Principal Component Analysis (PCA) based on my Global25 data. Levanluhta_IA are the Saami-related samples, while Levanluhta_IA_o is an Scandinavian-like outlier. Baltic_IA is an Iron Age individual from what is now Lithuania from the recent Damgaard et al. paper (see here). Note the accuracy of the Global25 data in pinpointing their genetic affinities and also the trajectory of the Levanluhta_IA cluster, which seems to be "pulling" towards Levanluhta_IA_o.
The Saami and Levanluhta_IA are clear outliers from the main Northern European cluster. There are two reasons for this: excess East Asian/Siberian-related ancestry and Saami-specific genetic drift. However, this eastern admixture and genetic drift are shared in varying degrees by other North European populations, especially those that also speak Uralic languages, and this is why they appear to be "pulling" towards the Saami/Levanluhta_IA clusters in my PCA. Thus, what this suggests is that the expansion of Uralic languages across Northeastern Europe was intimately linked with the spread of Siberian-related ancestry into the region.
This idea has been around for a long time and is now becoming even more widely accepted (see here). However, Lamnidis et al. also featured samples from a likely pre-Uralic (1523±87 calBCE) burial site at Bolshoy Oleni Ostrov in the Kola Peninsula, present-day northern Russia, and, perhaps surprisingly, found that they showed even more Siberian-related ancestry than Levanluhta_IA. So what's going on?
I'm confident that this discrepancy can be explained by multiple waves of migrations from the east into Northeastern Europe, possibly before, during and after the time of the people buried at Bolshoy Oleni Ostrov, by pre-Uralic, para-Uralic and/or Proto-Uralic-speaking populations.
Consider the following qpAdm output, in which Levanluhta_IA is just barely modeled successfully as a two-way mixture between Levanluhta_IA_o and Bolshoy_Oleni_Ostrov. The statistical fit improves significantly with the addition of Glazkovo_EBA as a third mixture source. This is an ancient population from near Lake Baikal dated to 4597-3726 BC from the aforementioned Damgaard et al. paper.
Levanluhta_IA Bolshoy_Oleni_Ostrov 0.468±0.036 Levanluhta_IA_o 0.532±0.036 chisq 19.129 tail prob 0.0854706 Full output Levanluhta_IA Bolshoy_Oleni_Ostrov 0.241±0.092 Glazkovo_EBA 0.162±0.059 Levanluhta_IA_o 0.597±0.046 chisq 7.756 tail prob 0.734966 Full outputFor the sake of being complete, I also tested whether Levanluhta_IA_o could be substituted by other similar ancient samples from the neighborhood, including those associated with the Battle-Axe and Corded Ware cultures. There's not much to report; qpAdm returned poor statistical fits and/or implausible ancestry proportions (for the full output from my runs, see here). Baltic_IA did produce a statistically sound model, but with excess Glazkovo_EBA-related ancestry. I also had to drop Bolshoy_Oleni_Ostrov from the analysis to make things work, which suggests to me that the result shouldn't be taken too literally.
Levanluhta_IA Baltic_IA 0.677±0.034 Glazkovo_EBA 0.323±0.034 chisq 8.547 tail prob 0.741095 Full outputSo as far as I can see, the western ancestry in Levanluhta_IA is likely to be mostly of Germanic origin, and thus Indo-European, meaning that it's logical to look east, perhaps far to the east, for the source of its Uralic ancestry. This might seem like a complicated and uncertain task, considering that Levanluhta_IA could well be at least a thousand years younger than the first entry of Uralic speakers into Fennoscandia. However, take a look what happens when I substitute Glazkovo_EBA with a variety of Uralic-speaking populations from around the Ural Mountains, which is where the Proto-Uralic homeland is generally considered to have been located.
Levanluhta_IA Bolshoy_Oleni_Ostrov 0.210±0.091 Khanty 0.283±0.090 Levanluhta_IA_o 0.507±0.035 chisq 7.007 tail prob 0.798532 Full output Levanluhta_IA Bolshoy_Oleni_Ostrov 0.193±0.098 Levanluhta_IA_o 0.495±0.035 Mansi 0.312±0.100 chisq 7.884 tail prob 0.7237 Full output Levanluhta_IA Bolshoy_Oleni_Ostrov 0.300±0.065 Levanluhta_IA_o 0.337±0.072 Mari 0.363±0.121 chisq 8.393 tail prob 0.677705 Full output Levanluhta_IA Bolshoy_Oleni_Ostrov 0.238±0.084 Levanluhta_IA_o 0.553±0.036 Nenets 0.209±0.067 chisq 7.210 tail prob 0.78181 Full output Levanluhta_IA Bolshoy_Oleni_Ostrov 0.302±0.069 Levanluhta_IA_o 0.324±0.081 Udmurt 0.373±0.135 chisq 9.195 tail prob 0.60393 Full outputAll of these models look great, and easily rival the best model with Glazkovo_EBA. Moreover, they make good sense in terms of linguistics. The only problem is that they're anachronistic, because the Uralic-speaking reference populations are younger than Levanluhta_IA. So I can't be certain that they reflect reality without corroboration from ancient DNA. It might turn out, for instance, that a Glazkovo_EBA-like population was already present somewhere deep in Europe before or during the time of Bolshoy_Oleni_Ostrov, while no such population existed around the Ural Mountains until the time of Levanluhta_IA. By the way, it might be important to note that the present-day Finnish samples in my dataset can't be modeled as a mixture between Levanluhta_IA and Levanluhta_IA_o. But they can be modeled as a mixture between Baltic_IA and Levanluhta_IA. I don't know which part of Finland they're from exactly; probably all over the place, so it'd be useful to test regional Finnish populations to see how they behave in such models. Of course, Finns aren't Saamic speakers, they're Finnic speakers, and they're probably the result of a more recent Uralic expansion into Fennoscandia than the one that gave rise to the Saami.
Finnish Baltic_IA 0.671±0.076 Levanluhta_IA 0.329±0.076 chisq 14.114 tail prob 0.293508 Full outputDamgaard et al. didn't report the Y-haplogroup for Baltic_IA, but the word round the campfire is that this individual belonged to N1c, which is today the most common Y-haplogroup among Uralic speakers. Obviously, we need a lot more ancient DNA to sort all of this out, but things are already looking pretty much as expected. Stay tuned for new posts in this series following the publication of more ancient DNA relevant to this fascinating topic. See also... How did Y-haplogroup N1c get to Bolshoy Oleni Ostrov? The Uralic cline in the Global25 Late PIE ground zero now obvious; location of PIE homeland still uncertain, but...
Labels:
ancient DNA,
Bolshoy Oleni Ostrov,
Corded Ware Culture,
CWC,
Fennoscandia,
Finnic,
Finno-Ugric,
Indo-European,
Levanluhta,
N-L1026,
N1c,
Northern Europe,
Proto-Uralic,
R1a-Z645,
Saami,
Siberia,
Uralic,
Urals
Thursday, November 1, 2018
Big deal of 2018: Yamnaya not related to Maykop
I was going to write this post after the genotype data from the Wang et al. preprint on the genetic prehistory of the Greater Caucasus became available, because I wanted to demonstrate a few key points with analyses of my own. But I've got a hunch that the formal publication of the manuscript, and thus also the release of the data, has been indefinitely delayed for one reason or another. So here goes anyway, the big deal of 2018...
This year, ancient DNA has revealed that the populations associated with the Maykop and Yamnaya archeological cultures were genetically distinct from each other, and, in all likelihood, didn't mix to any significant degree. Case in point: an ADMIXTURE analysis from Wang et al. 2018.
No doubt, this is quite a shock for many people, especially those of you who consider Maykop to have been a Proto-Indo-European-speaking culture that either gave rise to Yamnaya or at least Indo-Europeanized it. So now, if you still want to see Maykop as the Indo-Europeanizing agent in the Pontic-Caspian steppe, you'll have to rely solely on archeological and linguistics data, and also keep in mind that ancient DNA has slapped you in the face.
In just a few years, ancient DNA has provided us with plenty of shocks, but this is arguably among the biggest.
However, I honestly can't say that it was a huge surprise for me, because I tentatively predicted this outcome more than two years ago based on a handful of mitochondrial (mtDNA) haplotypes (see here). Certainly, analyzing genome-wide genetic data is what I thrive on, but if that's off limits, then eyeballing even a few mtDNA markers can also be very useful.
Wang et al. easily demonstrate the lack of any meaningful genetic relationship between Maykop (including Steppe Maykop, which shows an unusual eastern influence) and Yamnaya using a range of methods. But, judging by their conclusion, in which they still seem to want to see Maykop as the said Indo-Europeanizing agent in the Pontic-Caspian steppe, they're not exactly enthused by their own results. And they also make the following claim (emphasis is mine):
Based on PCA and ADMIXTURE plots we observe two distinct genetic clusters: one cluster falls with previously published ancient individuals from the West Eurasian steppe (hence termed ‘Steppe’), and the second clusters with present-day southern Caucasian populations and ancient Bronze Age individuals from today’s Armenia (henceforth called ‘Caucasus’), while a few individuals take on intermediate positions between the two. The stark distinction seen in our temporal transect is also visible in the Y-chromosome haplogroup distribution, with R1/R1b1 and Q1a2 types in the Steppe and L, J, and G2 types in the Caucasus cluster (Fig. 3A, Supplementary Data 1). In contrast, the mitochondrial haplogroup distribution is more diverse and almost identical in both groups (Fig. 3B, Supplementary Data 1).I'd say that what they're almost suggesting there is that the Caucasus and Steppe clusters, hence also the Maykop and Yamnaya populations, shared significant maternal ancestry. If this were true, then perhaps it might mean that the Pontic-Caspian steppe was Indo-Europeanized via female-biased migrations from Maykop? Yes, perhaps, if this were true. However, it's not. To be sure, Yamnaya does show a close genome-wide genetic relationship with an earlier group from the North Caucasus region: the so called Eneolithic steppe people. But they can't be linked to Maykop or even the roughly contemporaneous nearby Eneolithic Caucasus population, and seem to have vanished, at least as a coherent genetic unit, just as Maykop got going. Wang et al. managed to sequence three Eneolithic steppe samples with the following mtDNA haplogroups: H2, I3a and T2a1b. H2 is too broad a haplogroup to bother with, but here are the results for I3a and T2a1b from the recently launched AmtDB, the first database of ancient human mitochondrial genomes (see here). In a database of 1,131 ancient samples, I3a shows up in just five individuals, all of them associated with Yamnaya-related archeological cultures and populations: Poltavka (BARu), Unetice (UNC), Corded Ware (CWC), and Bell Beaker (BBC). Similarly, T2a1b shows up in just four individuals, all of them associated with Corded Ware (CWC) and Bell Beaker-derived Bronze Age Britons (BABI). And if I go back a step to T2a1, then the list reveals two Yamnaya individuals from what is now Kalmykia, Russia. Thus, using just two mtDNA haplotypes I'm able to corroborate the results from genome-wide genetic data showing a close relationship between Eneolithic steppe and Yamnaya. So like I said, useful stuff. This obviously begs the question: what does the AmtDB reveal about Maykop mtDNA haplotypes, especially in the context of the genetic relationship, or rather lack of, between Yamnaya and Maykop? Yep, again, the AmtDB basically corroborates the results from genome-wide genetic data. But don't take my word for it. Stick the currently available Maykop mtDNA haplogroups into the AmtDB and see what happens (for your convenience I've made a list available here). Considering the close geographic and temporal proximity of Maykop to Yamnaya, you won't see an overly high sharing rate with Yamnaya and closely related populations. Moreover, Maykop shows several haplogroups that appear highly unusual in the context of the Eneolithic and Bronze Age steppe mtDNA gene pool, and, instead, link its maternal ancestry to those of the early European farmers, West Asians or even Central Asians, such as HV, M52, U1b, U7b and X2f. See also... Yamnaya: home-grown Yamnaya isn't from Iran just like R1a isn't from India Big deal of 2016: the territory of present-day Iran cannot be the Indo-European homeland
Saturday, September 22, 2018
Corded Ware people =/= Proto-Uralics (Tambets et al. 2018)
A new paper on the genetic structure of Uralic-speaking populations has appeared at Genome Biology (see here). It looks to me like the prelude to a forthcoming paleogenetics paper on the same topic that was discussed in the Estonian media recently (see here). Although not exactly ground breaking (because it basically argues what I've been saying at this blog for years, like here), it's a very nice effort all round and must be read by anyone with an interest in this topic. From the paper, emphasis is mine:
Background The genetic origins of Uralic speakers from across a vast territory in the temperate zone of North Eurasia have remained elusive. Previous studies have shown contrasting proportions of Eastern and Western Eurasian ancestry in their mitochondrial and Y chromosomal gene pools. While the maternal lineages reflect by and large the geographic background of a given Uralic-speaking population, the frequency of Y chromosomes of Eastern Eurasian origin is distinctively high among European Uralic speakers. The autosomal variation of Uralic speakers, however, has not yet been studied comprehensively. Results: Here, we present a genome-wide analysis of 15 Uralic-speaking populations which cover all main groups of the linguistic family. We show that contemporary Uralic speakers are genetically very similar to their local geographical neighbours. However, when studying relationships among geographically distant populations, we find that most of the Uralic speakers and some of their neighbours share a genetic component of possibly Siberian origin. Additionally, we show that most Uralic speakers share significantly more genomic segments identity-by-descent with each other than with geographically equidistant speakers of other languages. We find that correlated genome-wide genetic and lexical distances among Uralic speakers suggest co-dispersion of genes and languages. Yet, we do not find long-range genetic ties between Estonians and Hungarians with their linguistic sisters that would distinguish them from their non-Uralic-speaking neighbours. Conclusions: We show that most Uralic speakers share a distinct ancestry component of likely Siberian origin, which suggests that the spread of Uralic languages involved at least some demic component. ... Recent aDNA studies have shown that extant European populations draw ancestry form three main migration waves during the Upper Palaeolithic, the Neolithic and Early Bronze Age [2, 3, 45]. The more detailed reconstructions concerning NE Europe up to the Corded Ware culture agree broadly with this scenario and reveal regional differences [65–67]. However, to explain the demographic history of extant NE European populations, we need to invoke a novel genetic component in Europe—the Siberian. The geographic distribution of the main part of this component is likely associated with the spread of Uralic speakers but gene flow from Siberian sources in historic and modern Uralic speakers has been more complex, as revealed also by a recent study of ancient DNA from Fennoscandia and Northwest Russia [68]. Thus, the Siberian component we introduce here is not the perfect but still the current best candidate for the genetic counterpart in the spread of Uralic languages.Citation... Tambets et al., Genes reveal traces of common recent demographic history for most of the Uralic-speaking populations, Genome Biology, (2018) 19:139 https://doi.org/10.1186/s13059-018-1522-1 See also... Big deal of 2019: ancient DNA confirms the link between Y-haplogroup N and Uralic expansions
Subscribe to:
Posts (Atom)